A Long-Context Generative Foundation Model Deciphers RNA Design Principles
A Long-Context Generative Foundation Model Deciphers RNA Design Principles
Huang, Y.; Lv, G.; Cheng, A.; Xie, W.; Chen, M.; Ma, X.; Huang, Y.; Tang, Y.; Shi, Q.; Wang, Z.; Wang, J.; Yunpeng, X.; Zhao, L.; Cai, Y.; Chen, J. X.; Zheng, S.
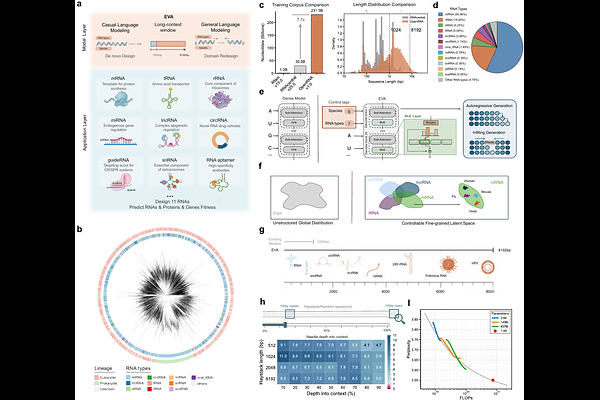
AbstractProgrammable design of RNA sequences with defined functions remains a central challenge in biology. Despite recent advances, existing RNA generative models lack robust controllable design capabilities and are constrained by short context windows, limiting their capacity to model the complex evolutionary manifold of full-length transcripts. Here, we introduce EVA (Evolutionary Versatile Architect), a long-context generative RNA foundation model trained on over 114 million full-length RNA sequences spanning the full breadth of evolutionary diversity. By integrating a Mixture-of-Experts architecture with an 8,192-token context window, EVA learns a unified, evolution-consistent representation of RNA sequence space that captures both global transcript structure and fine-grained functional elements. EVA establishes a unified framework for a broad spectrum of RNA design tasks within a single architecture, ranging from mutational fitness prediction to context-aware sequence engineering. We demonstrate EVA's versatility through the de novo design and targeted optimization of diverse RNA classes, including tRNAs, aptamers, and CRISPR guide RNAs, as well as mRNA and circular RNA therapeutics. In systematic evaluations, EVA achieves state-of-the-art performance across seven of nine established benchmarks, improving structural modeling accuracy by up to an order of magnitude compared to existing approaches. EVA is fully open-source at the following repository: https://github.com/GENTEL-lab/EVA.