Processing of Reversed Replication Forks is Required for the Resolution of Replication-Transcription Conflicts
Processing of Reversed Replication Forks is Required for the Resolution of Replication-Transcription Conflicts
Carvajal-Garcia, J.; Merrikh, H.
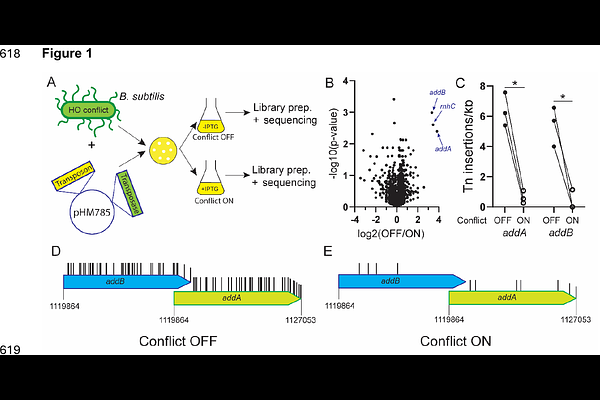
AbstractDNA replication and transcription occur simultaneously on the same template, leading to conflicts between the two machineries. Conflicts stall replication forks, lead to genome instability, mutagenesis, and the formation of deleterious R-loop structures. There are two types of conflicts: depending on the strand where a gene is encoded, the two machineries either meet head-on or co-directionally. The adverse outcomes of conflicts in the head-on orientation are significantly more detrimental to cells compared to co-directional conflicts. Despite many studies across various organisms, how the replication fork structure is impacted by these encounters remains unclear. Here, we performed an unbiased genetic screen using a transposon library to identify factors essential for surviving head-on conflicts in Bacillus subtilis. Our screen identified three hits: RNase HIII, AddA and AddB. Our prior work had shown that RNase HIII, which processes R-loops, is essential for surviving head-on conflicts. However, AddA and AddB, which function together as a complex to process blunt DNA ends, have not been previously identified as essential conflict resolution factors. Through follow-up genetic and biochemical analyses, we found that the helicase activity but not the nuclease activity of this complex is required for conflict resolution. Based on the fundamental properties of DNA at replication fork structures, our work collectively indicates that upon head-on conflicts, the nascent strands form a reversed fork structure, which is then unwound by AddAB, which leads to re-annealing to the parental strands. This process re-establishes an intact replication fork that can be used to restart replication after conflicts with transcription.