Beyond gene length: Exon-intron architecture and isoform potential in the evolution of eukaryotic complexity
Beyond gene length: Exon-intron architecture and isoform potential in the evolution of eukaryotic complexity
Lu, S.; Bao, Y.; Sheynkman, G. M.; Korkin, D.
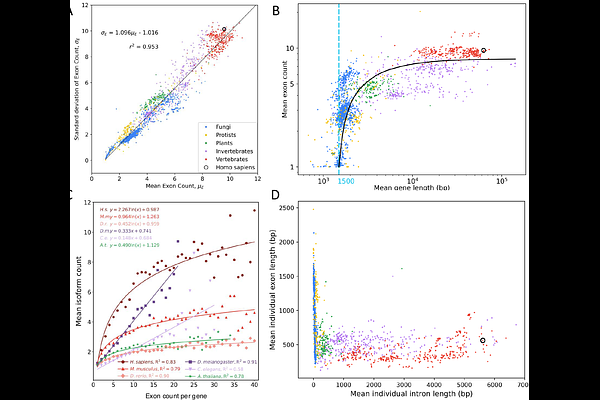
AbstractAlternative splicing is a major source of human transcriptomic and phenotypic variation, yet its evolutionary contribution to genomic complexity remains unresolved. It has been shown that a mean gene length can be the most basic but remarkably efficient proxy for multicellular genome complexity, whereas mean protein length is not, as it plateaus abruptly early in eukaryotic evolution. Here, we show that, across 2,683 genomes, exon count continues to increase beyond this transition and then rapidly saturates at ~10 exons per gene, supporting its role as an additional dimension of genomic complexity linked to exon-intron architecture. A simple stochastic exon-splitting model reproduces the observed biphasic exon-number growth pattern and identifies minimal exon length as a key determinant.