23S rRNA modifications stimulate catalytic activity and prevent the formation of alternative structures
23S rRNA modifications stimulate catalytic activity and prevent the formation of alternative structures
Larsson, D. S. D.; Liiv, A.; Ero, R.; Remme, J.; Selmer, M.
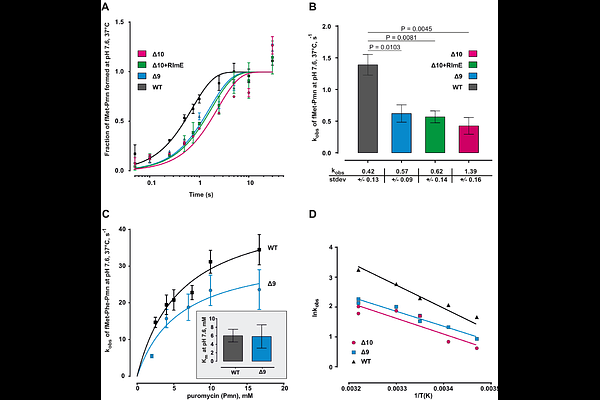
AbstractRibosomal RNA modifications cluster around the peptidyl transferase center (PTC), the catalytic center of the ribosome, yet their collective functional roles remain unclear. Here we analyze Escherichia coli ribosomes lacking 11 or 12 modifications near the PTC. Using kinetic assays, we show these hypo-modified ribosomes catalyze peptide bond formation at rates two- to threefold lower than wild-type and exhibit reduced thermal stability. Cryo-electron microscopy of hypo-modified ribosomes reveals multiple alternative conformations of the PTC and exit tunnel regions, disrupting native stacking and hydrogen bonding critical for positioning of tRNA substrates. These findings indicate that rRNA modifications stabilize the native PTC structure, preventing formation of alternative, nonfunctional conformations and thereby enhance catalytic efficiency. Our study provides insight into how rRNA modifications fine-tune ribosome function by maintaining structural integrity essential for efficient translation.