Nonlinear mixed-effect models and tailored parametrization schemes enables integration of single cell and bulk data
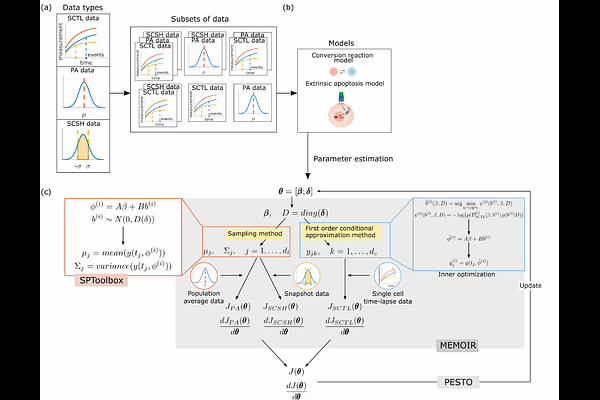
Nonlinear mixed-effect models and tailored parametrization schemes enables integration of single cell and bulk data
Wang, D.; Froehlich, F.; Stapor, P.; Schaelte, Y.; Huth, M.; Eils, R.; Kallenberger, S.; Hasenauer, J.
AbstractExperimental methods for characterizing single cells and cell populations have improved tremendously over the past decades. This progress has enabled the development of quantitative, mechanistic models for cellular processes based on either single cell or bulk data. However, coherent statistical frameworks for the model-based integration of different data types at the single-cell and population levels are still missing. In this work, we present a mathematical modeling approach for integrating single-cell time-lapse, single-cell snapshot, single-cell time-to-event and population-average data. Utilizing a formulation based on nonlinear mixed-effect modeling, we enable the description of multiple data types, with and without single-cell resolution, and we propose a tailored parameter estimation method. Furthermore, we propose a tailored parameter estimation scheme that facilitates the assessment of underlying process parameters. Our study demonstrates that the proposed approach can reliably integrate diverse data types, thereby improving parameter identifiability and prediction accuracy. Applying this framework of extrinsic apoptosis reveals that simultaneously considering multiple data types can be essential, particularly when experimental constraints limit data availability. The proposed approach is broadly applicable and may significantly advance our understanding of complex biological processes.