Spatial Mechanomics for Tissue-Scale Biomechanical Mapping and Multi-omics Integration
Spatial Mechanomics for Tissue-Scale Biomechanical Mapping and Multi-omics Integration
Xie, W.; Wang, Z.; Shan, Q.; Zhao, Q.; Ye, X.
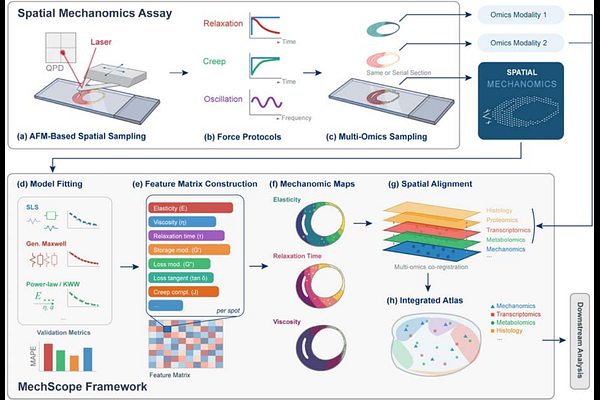
AbstractTissue mechanical properties are spatially heterogeneous and tightly coupled to cellular function, developmental patterning, and disease progression, yet spatially resolved characterization of viscoelastic and microrheological behavior across intact tissues remains limited. Here we introduce spatial mechanomics, a framework for tissue-wide acquisition, quantitative extraction, and computational representation of location-resolved mechanical states. Using BioAFM-based spatial sampling with multi-protocol microrheology, we acquire force responses at defined tissue coordinates and fit physically interpretable viscoelastic models to extract elastic, viscous, and frequency-dependent parameters at each position. These parameters are assembled into per-niche mechanomic feature vectors and reconstructed into tissue-scale mechanomic atlases that resolve heterogeneous mechanical organization. We implement these capabilities in MechScape, an open-source computational platform that supports force curve fitting, spatial feature matrix construction, unsupervised domain discovery, and cross-modal alignment with histological and molecular measurements. Application to murine myocardial tissue reveals that spatial mechanomics identifies distinct mechanical states, quantifies condition-dependent remodeling across all measured parameters, and resolves spatially coherent mechanical domains. This work establishes spatial mechanomics as a quantitative approach for tissue-scale biomechanical mapping and provides a generalizable framework for integrating mechanics as an omics layer in multi-modal tissue analysis.