Expression landscape of heterologous enzymes in Synechocystis sp. PCC 6803
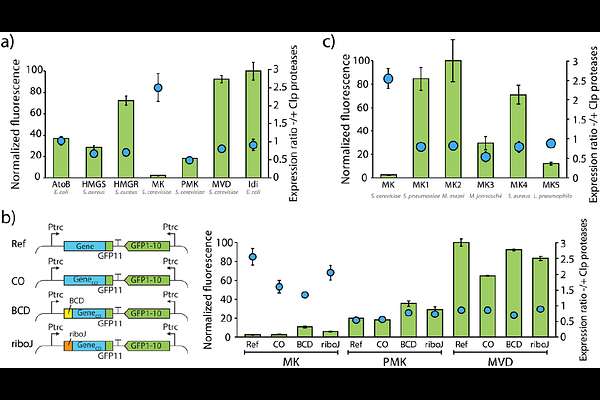
Expression landscape of heterologous enzymes in Synechocystis sp. PCC 6803
Medipally, H.; Karlsson, A.; Dheer, A.; Hudson, E. P.; Englund, E.
AbstractPhotosynthetic cyanobacteria are promising platforms for sustainable chemical production, as they can convert light and CO2 into valuable compounds. Achieving this often requires engineering cyanobacteria with non-native enzymes with strong promoters to maximize enzyme accumulation. However, despite extensive engineering efforts, the extent to which heterologous proteins misfold and undergo degradation in cyanobacteria remains unknown. Here, we systematically investigate the fate of recombinant proteins in Synechocystis sp. PCC 6803 by quantifying metabolic enzyme degradation. To do this, we developed a quantitative approach that combines split-GFP protein reporting with inducible CRISPRi knockdown of Clp protease system, enabling detection of proteins that would otherwise be degraded. Applying this method to 103 heterologous proteins previously used in cyanobacterial metabolic engineering studies, we find that nearly half undergo significant degradation, with some losing over 95% of their potential expression. Furthermore, we demonstrate that replacing enzymes with homologs is often a more effective strategy to address expression issues than optimizing genetic elements. These findings provide the first quantitative overview of heterologous protein expression in cyanobacteria and identify enzymes that are poorly expressed and suboptimal for their respective pathways, information usable to increase production titers in photosynthetic cell factories.