Benchmarking ambient RNA removal across droplet and well-plate platforms reveals artificial count generation as a critical failure mode of scAR and CellClear
Benchmarking ambient RNA removal across droplet and well-plate platforms reveals artificial count generation as a critical failure mode of scAR and CellClear
Schroeder, L.; Gerber, S.; Ruffini, N.
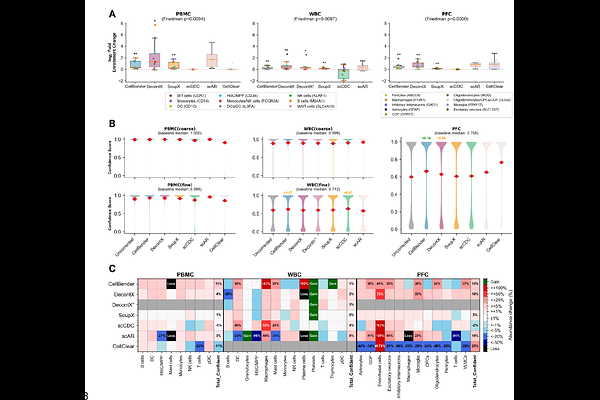
AbstractBackground: Ambient RNA contamination is a pervasive artifact of single-cell and single-nucleus RNA sequencing (sxRNA-seq), yet no consensus exists on which computational removal tool performs best across experimental platforms. Results: We present a systematic benchmark of six tools: CellBender, DecontX, SoupX, scCDC, scAR, and CellClear - evaluated across six human-mouse cell line mixing (hgmm) datasets (1k-20k cells) providing partial ground truth, two droplet-based complex tissue datasets (PBMC scRNA-seq; prefrontal cortex snRNA-seq), and a well-plate-based dataset (BD Rhapsody WBC). Using inter-species counts as partial ground truth, we quantify sensitivity, specificity, precision, and removal consistency per tool. We further apply a count-integrity criterion quantifying gene-cell positions where corrected values exceed raw counts. This reveals that scAR and CellClear do not merely denoise but fundamentally restructure count matrices: CellClear replaces >93% of counts with values derived from matrix factorization, while scAR generates spurious cell types absent from uncorrected data, including three spurious coarse cell types in the BD Rhapsody dataset and up to eight novel cell types in the prefrontal cortex. CellBender and SoupX exhibit reliable contamination removal with minimal count distortion. DecontX and scCDC are the only tools operable on non-droplet platforms without raw count matrix access. Runtime benchmarking at atlas scale (up to 172,000 nuclei) further demonstrates that CellClear fails to scale. Conclusions: Count matrix integrity, not removal sensitivity alone, must be a primary criterion when selecting ambient RNA correction tools. We provide platform-specific recommendations and a decision framework to guide tool selection across experimental contexts.