Learning latent conformational landscapes encoded in cryo-EM
Learning latent conformational landscapes encoded in cryo-EM
Dai, H.; Shen, Y.; Chen, Q.; Li, L.; Xu, Z.; Li, M.; Xie, Y.; Zheng, J.; Pei, Y.; Zhang, J.; Sun, L.; Liu, Z. J.; Yu, J.
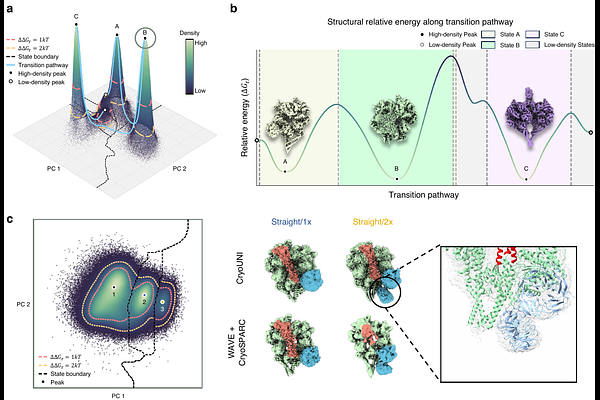
AbstractProteins exist as a continuum of conformational states, millions of snapshots of which are captured in every cryo-EM experiment. Current analysis reduces these snapshots to static density maps or discrete classes, discarding the dynamics encoded in the data. Continuous latent-space methods offer a promising alternative, yet whether their learned representations are physically grounded remains unresolved. Here, we show that cryo-EM data can be organized into a probabilistic conformational landscape in which latent density reflects the distribution of molecular states. We demonstrate that this landscape is both physically grounded and experimentally actionable. For integrin v{beta}8, the learned landscape shows strong agreement with independently derived molecular dynamics simulations, supporting its physical plausibility. Additionally, the landscape reveals biologically meaningful organization of molecular states, including a spectrum of states in LIS1-mediated dynein activation, from dominant conformations to low-population intermediates defined by distinct binding stoichiometries, and continuous conformational pathways in KCTD5/CUL3NTD/G{beta}{gamma} complex that are consistent with prior simulations. Probability-guided particle selection further improves reconstruction quality, closing the loop from landscape to structure determination. These results establish that cryo-EM data encode a latent conformational landscape that can be directly exploited for structural reconstruction, discovery, and interpretation.