Kinome profiling allows examination and prediction of kinase inhibitor cardiotoxicity
Kinome profiling allows examination and prediction of kinase inhibitor cardiotoxicity
Tabet, J. S.; Joisa, C. U.; Jensen, B. C.; Gomez, S. M.
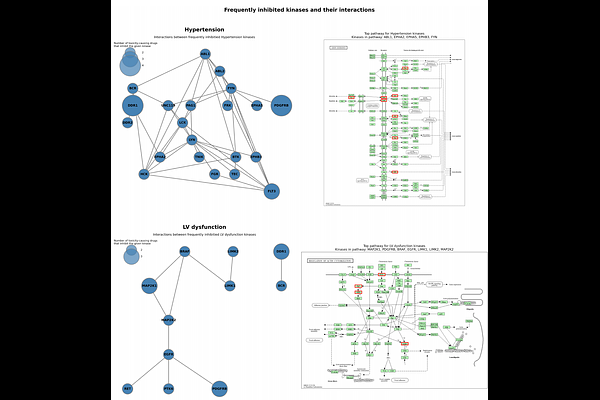
AbstractBackground: Despite improved cancer outcomes with kinase inhibitors (KIs), their cardiotoxicity remains a significant clinical challenge. Current approaches to predict and prevent KI-induced cardiac adverse events (CAEs) are limited by an incomplete understanding of underlying mechanisms, including the contribution of off-target kinase engagement. Objectives: To establish links between kinase inhibition profiles and cardiotoxic phenotypes using empirical proteomic data, and to leverage these profiles in machine learning (ML) models capable of predicting KI cardiotoxicity. Methods: We curated a database connecting kinome-wide target binding profiles of FDA-approved KIs (n=44) with documented incidence rates of six distinct CAEs. Binding profiles were derived from unbiased chemoproteomics and used to assess associations between KI selectivity, specific kinase targets, and CAEs. Profiles were further used to develop ML models to predict CAE risk, with SHAP-based model interpretation applied to identify cardiotoxicity-associated kinases. Results: KI promiscuity was not a significant predictor of cardiotoxicity across all six CAEs. Frequency analysis revealed that kinases including RET, PDGFRB, and DDR1 are recurrently inhibited across CAE-linked compounds, with nearly all identified as off-targets not annotated by the FDA. Network and pathway enrichment analyses supported a systems-level model in which cardiotoxicity arises from coordinated disruption of cardiac-relevant signaling networks. ML models achieved 66-84% cross-validated accuracy (ROC-AUC 0.75-0.8) across CAE endpoints, with SHAP analysis identifying PDGFRB, EGFR, and MEK1/2 among the most predictive kinases. Conclusions: Proteomic kinome profiling combined with machine learning provides a mechanistically grounded framework for predicting KI cardiotoxicity and supports off-target-aware drug design to minimize cardiovascular risk.