PepBERT: Lightweight language models for peptide representation
PepBERT: Lightweight language models for peptide representation
Du, Z.; Li, Y.
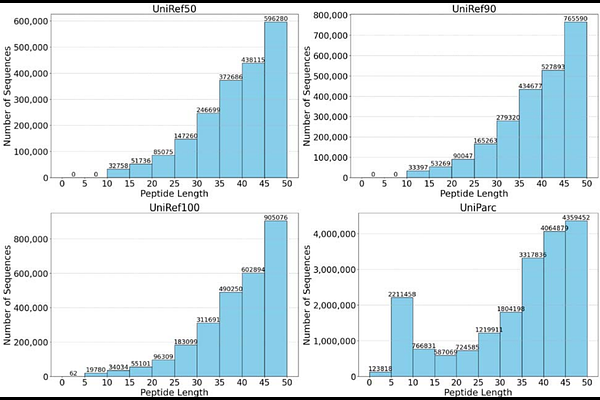
AbstractProtein language models (pLMs) have been widely adopted for various protein and peptide-related downstream tasks and demonstrated promising performance. However, short peptides are significantly underrepresented in commonly used pLM training datasets. For example, only 2.8% of sequences in the UniProt Reference Cluster (UniRef) contain fewer than 50 residues, which potentially limit the effectiveness of pLMs for peptide-specific applications. Here, we present PepBERT, a lightweight and efficient peptide language model specifically designed for encoding peptide sequences. Two versions of the model, PepBERT-large (4.9 million parameters) and PepBERT-small (1.86 million parameters), were pretrained from scratch using four custom peptide datasets and evaluated on nine peptide-related downstream prediction tasks. Both PepBERT models achieved performance superior or comparable to the benchmark model, ESM-2 with 7.5 million parameters, on 8 out of 9 datasets. Overall, PepBERT provides a compact yet effective solution for generating high-quality peptide representations for downstream applications such as bioactive peptide screening and drug discovery. The datasets, source codes, pretrained models, and tutorials for usage of PepBERT are available at https://github.com/dzjxzyd/PepBERT-large.