AlphaFold Database expands to proteome-scale quaternary structures
AlphaFold Database expands to proteome-scale quaternary structures
Han, Y.; Tsenkov, M. I.; Venanzi, N. A. E.; Bertoni, D.; Cha, S.; Chacon, A.; Dietrich, N.; Fomitchev, B.; Goldtzvik, Y.; Hsu, D.; Austin, J.; Ellaway, J.; Didi, K.; Kovalevskiy, O.; Lasecki, D.; Laydon, A.; Livne, M.; Magana, P.; Majewski, M.; Nair, S.; Paramval, U.; Patel, N.; Patel, R.; Pidruchna, I.; Santini Lopez, B.; Sohani, P.; Tanweer, A.; Tran, D.; Tretina, K.; Vollmar, M.; Vu, Q.; Zidek, A.; Velankar, S.; Steinegger, M.; Fleming, J.; Mirdita, M.; Dallago, C.
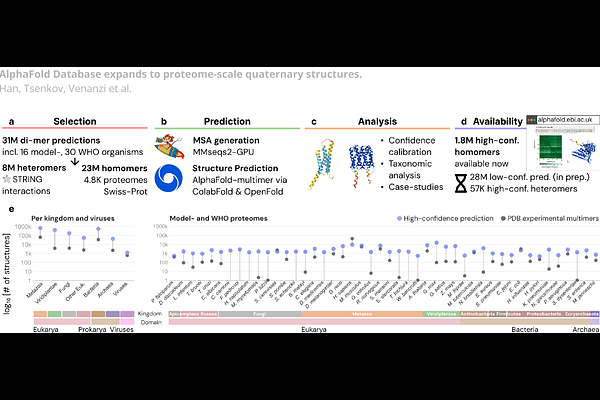
AbstractProtein function is governed by molecular interactions, yet structural coverage of these interactions remains sparse. The AlphaFold Protein Structure Database (AFDB) transformed access to accurate monomeric protein structures at scale. Here, we expand AFDB with 1.8M high-confidence protein complexes by conducting a large-scale study of over 31M predicted homo- and heteromeric protein complexes compiled from 4,777 proteomes, including model- and global health organisms, and using STRING physical-interaction annotations. We calibrate confidence metrics to assess the quality of complex predictions, and propose confidence cutoffs. These enabled the discovery of emergent structure and topologies in complex structure prediction that is not present with monomeric predictions. Clustering of high-confidence complexes showed that the largest 1% of non-singleton representatives account for ~25% of all complexes, and that ~9% of clusters are conserved across superkingdoms. In summary, large-scale structural predictions of the interactome serve as a foundational resource to facilitate functional and mechanistic discovery across biology.