Ancestral state reconstruction with discrete characters using deep learning
Ancestral state reconstruction with discrete characters using deep learning
Nagel, A. A.; Landis, M. J.
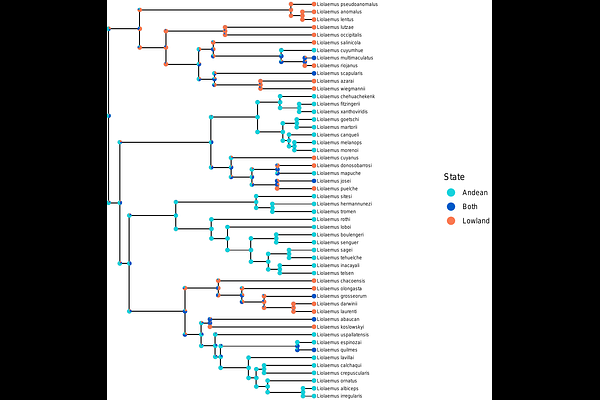
AbstractAncestral state reconstruction is a classical problem of broad relevance in phylogenetics. Likelihood-based methods for reconstructing ancestral states under discrete character models, such as Markov models, have proven extremely useful, but only work so long as the assumed model yields a tractable likelihood function. Unfortunately, extending a simple but tractable phylogenetic model to possess new, but biologically realistic, properties often results in an intractable likelihood, preventing its use in standard modeling tasks, including ancestral state reconstruction. The rapid advancement of deep learning offers a potential alternative to likelihood-based inference of ancestral states, particularly for models with intractable likelihoods. In this study, we modify the phylogenetic deep learning software phyddle to conduct ancestral state reconstruction. We evaluate phyddle's performance under various methodological and modeling conditions, while comparing to Bayesian inference when possible. For simple models and small trees, its performance resembles the performance of Bayesian inference, but worsens as tree size increases. While phyddle still performs adequately for more complex models, such as speciation and extinction models, the estimates differ more from Bayesian inference in comparison with simpler models. Lastly, we use phyddle to infer ancestral states for two empirical datasets, one of the ancestral ranges of a subclade of the genus Liolaemus and ancestral locations for sequences from the 2014 Sierra Leone Ebola virus disease outbreak.