Fung-AI: An AI/ML-driven pipeline for antifungal peptide discovery
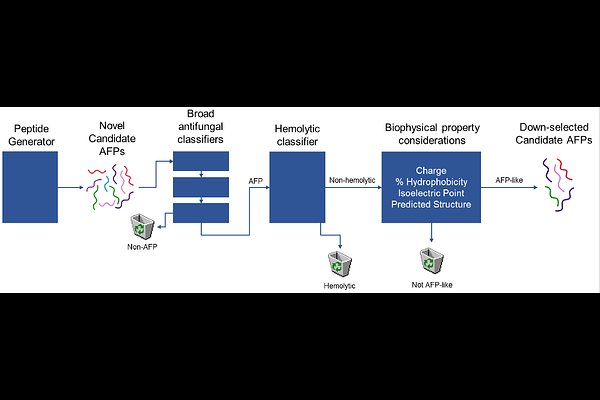
Fung-AI: An AI/ML-driven pipeline for antifungal peptide discovery
Berman, D. S.; Lewis, L. M.; Curtis, T. D.; Tiburzi, O. N.; Smith, D. F.; Casadevall, A.; Dunphy, L.
AbstractEmerging fungal pathogens represent a concerning threat to both global health and food security. In this study, we aimed to address our rising vulnerability to fungal pathogens through the development of the Fung-AI pipeline: an AI/ML-driven approach for antifungal discovery. A generative adversarial network (GAN) was trained to generate novel candidate antifungal peptide sequences. Next, in silico antifungal and hemolytic classifiers were built to further prioritize AI-generated peptides for experimental validation. From a pool of ~10,000 candidates, thirteen peptides were selected for testing over two-stages of experimentation. Five peptides were found to display mild antifungal activity against the wheat pathogen, Fusarium graminearum, with minimal inhibitory concentrations (MICs) ranging from 250 {micro}g/mL to 500 {micro}g/mL. Four of the five peptides also showed activity against the human pathogen, Candida albicans (MIC: 500 {micro}g/mL). Two of our AI-generated antifungal peptides additionally demonstrated low cytotoxicity in HepG2 human liver carcinoma cells (LC50 > 704.2 {micro}g/mL) indicating that they may be useful as scaffolds for future optimization for therapeutic applications. None of our peptides were found to considerably inhibit the emerging pathogen C. auris, suggesting the need for pathogen-specific down-selection of candidate peptides. Overall, we present a proof-of-principle, generative-AI-based approach for the rapid design of de novo antifungal peptides.