Structure-aware geometric graph learning for modeling protease-substrate specificity at scale
Structure-aware geometric graph learning for modeling protease-substrate specificity at scale
Guo, X.; Bi, Y.; Ran, Z.; Pan, T.; Sun, H.; Hao, Y.; Jia, R.; Wang, C.; Zhang, Q.; Kurgan, L.; Song, J.; Li, F.
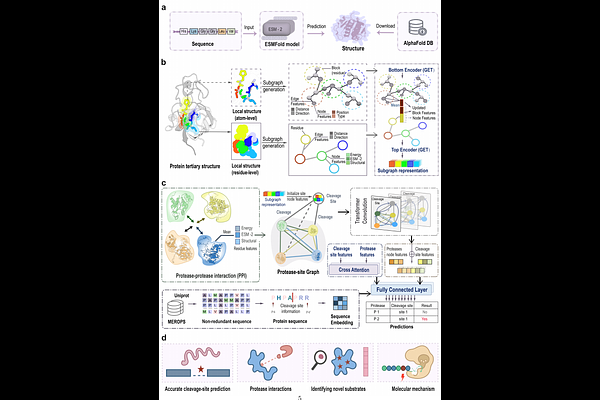
AbstractProtease-substrate specificity is central to cellular regulation and disease pathogenesis, and accurately modeling its structural determinants remains challenging. Substrate recognition is governed by spatial constraints and higher-order relationships that extend beyond local sequence motifs. Most computational approaches rely predominantly on motif-centric or sequence-based representations, limiting their ability to capture the geometric and relational structure underlying enzymatic specificity. Here, we introduce OmniCleave, a structure-aware geometric graph learning framework for modeling protease-substrate specificity at scale. OmniCleave is trained on 57,278 structure-informed protease-substrate pairs derived from 9,651 substrates spanning over 100 proteases across six distinct families. The framework integrates multi-scale structural graphs with higher-order protease relational topology, explicitly encoding spatial context and inter-protease dependencies within a unified geometric representation. This formulation moves beyond local pattern recognition and enables transferable modelling across six protease families. Across large-scale benchmarks, the framework consistently outperforms existing approaches and reveals interpretable geometric determinants underlying substrate recognition. Experimental validation confirms three novel caspase-3 substrates and 21 cleavage sites predicted by OmniCleave, supporting the biological relevance of the learned representations. Together, OmniCleave provides a scalable geometric framework for modeling protease-substrate specificity, with practical utility for systematic analysis of protease biology.