MAP: A Knowledge-driven Framework for Predicting Single-cell Responses for Unprofiled Drugs
MAP: A Knowledge-driven Framework for Predicting Single-cell Responses for Unprofiled Drugs
Feng, J.; Zhao, Z.; Zhang, X.; Liu, M.; Chen, J.; Quan, X.; Zhang, J.; Wang, Y.; Zhang, Y.; Xie, W.
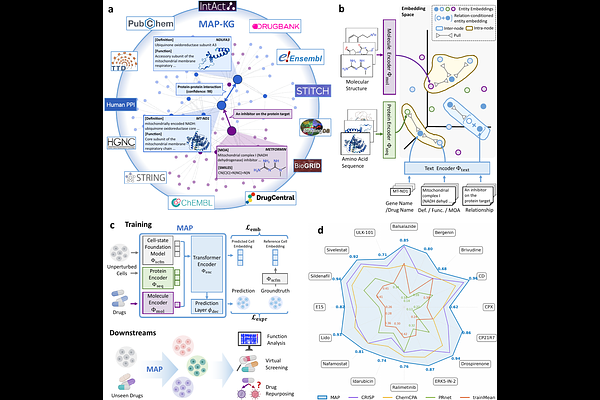
AbstractPredicting how cells respond to chemical perturbations is one of the goals for building virtual cells, yet experimentally profiled compounds cover only a small fraction of this space. Existing models struggle to generalize to unprofiled compounds, as they typically treat drugs as isolated identifiers without encoding their mechanistic relationships. We present MAP, a framework that integrates structured biological knowledge into cellular perturbation modeling and supports zero-shot prediction for small molecules with scarce or absent perturbation profiles. Specifically: (i) we construct MAP-KG, a large-scale knowledge graph tailored for cellular perturbation modeling that unifies 14 public resources, spanning 187k drugs, 23k genes, and 694k mechanistic relationships; (ii) we propose a knowledge-driven pre-training strategy that aligns molecular structures, protein sequence features, and textual mechanistic descriptions into a unified embedding space via contrastive learning, producing mechanism-aware and transferable gene and compound embeddings. The resulting knowledge-informed gene and drug representations are then coupled with a pretrained single-cell foundation model to condition perturbation response prediction; (iii) we evaluate MAP under two zero-shot generalization regimes: unseen cell type-drug combinations and the stricter setting of unprofiled drugs, where it improves top-50 DEG Pearson delta correlation by up to +13.3% and +12.2%, respectively, over the strongest baselines across three benchmarks. We further perform pathway-level functional analysis via GSEA for in-silico screening, where MAP predicts coherent, mechanism-consistent programs on unprofiled candidate drugs, and prioritizes 4 of 5 approved anti-cancer drugs in A-549 (non-small cell lung cancer).