Systematic CRISPRi screening reveals genetic modulators of E. coli isoprenoid production
Systematic CRISPRi screening reveals genetic modulators of E. coli isoprenoid production
Dokwal, D.; Brown, P. M.; Ingle, C.; Saunders, S. H.; Reynolds, K. A.
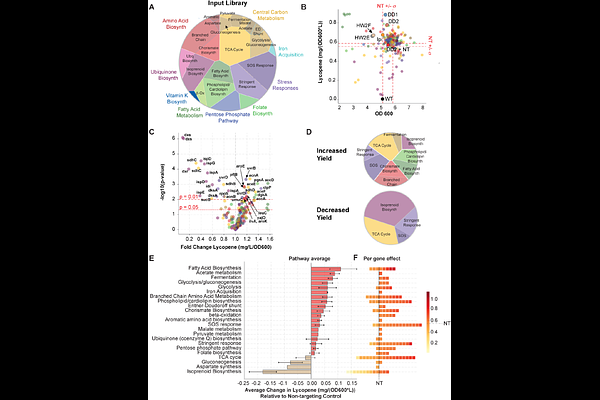
AbstractIsoprenoid biosynthesis in E. coli is a promising source of high-value natural products. However, isoprenoid production imposes a substantial metabolic burden on cellular carbon and cofactor metabolism. Optimizing yields thus requires rebalancing gene expression throughout metabolism. To systematically identify the genes in E. coli central metabolism shaping isoprenoid yield, we established a high-throughput CRISPR interference (CRISPRi) assay to examine the impact of gene repression on production of the bright red carotenoid, lycopene. Our approach enables repression of both essential and non-essential genes and links gene expression perturbations to pathway yield and biomass. We screened a CRISPRi library targeting 180 E. coli genes and found 31 genes for which repression significantly modified lycopene yield. Genes whose repression increased lycopene yield were found across fatty acid, amino acid, central carbon, and isoprenoid metabolism as well as stress response pathways. This set included four genes in amino acid biosynthesis, one in phospholipid biosynthesis, and one in the stringent response that, to our knowledge, have not previously been implicated in lycopene production. In contrast, genes for which repression decreased yield were limited to isoprenoid biosynthesis, central carbon metabolism, and stress response pathways. Together, our work reveals genetic targets for increasing lycopene yield throughout metabolism, and defines a tractable, generalizable approach to mapping genetic factors that modulate biosynthetic pathway yield.