Machine learning-based exploration, expansion and definition of the atropopeptide family of ribosomally synthesized and posttranslationally modified peptides
Machine learning-based exploration, expansion and definition of the atropopeptide family of ribosomally synthesized and posttranslationally modified peptides
Biermann, F.; Tan, B.; Breitenbach, M.; Nanudorn, P.; Dimitrova, Y.; Kakumu, Y.; Walker, A. S.; Ueoka, R.; Helfrich, E. J. N.
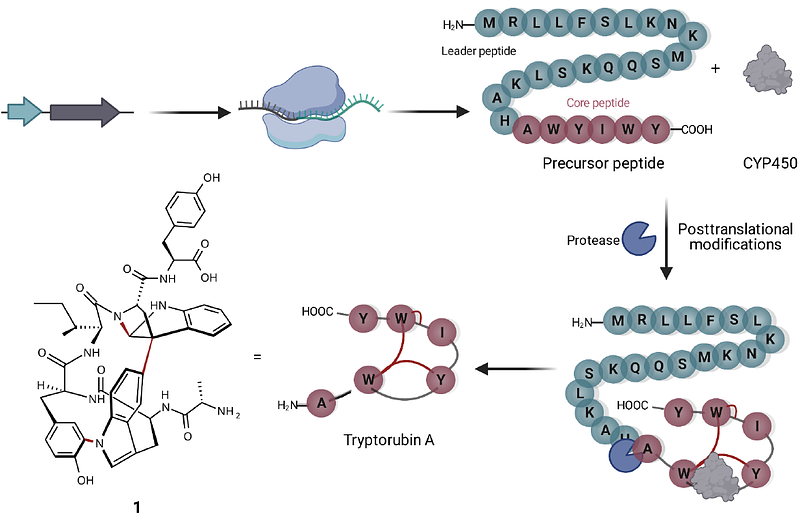
AbstractRibosomally synthesized and posttranslationally modified peptides (RiPPs) constitute a diverse class of natural products. Atropopeptides are a recent addition to the fast-growing number of RiPP families. Characterized members of the peptide family feature a particular intricate three-dimensional shape. Here we developed AtropoFinder, a machine learning-based algorithm to chart the biosynthetic landscape of the atropopeptides. AtropoFinder identified more than 650 atropopeptide biosynthetic gene clusters (BGCs). Through bioinformatics and modeling analyses, we pinpointed crucial motifs and residues in leader and core peptide sequences, prompting a refined definition of the atropopeptide RiPP family. Our study revealed that a substantial subset of atropopeptide BGCs harbors multiple tailoring genes, thus suggesting a broader structural diversity than previously anticipated. To verify AtropoFinder, we heterologously expressed four atropopeptide BGCs, which resulted in the identification of novel atropopeptides with varying peptide lengths, number and type of modifications. Most notably, our study resulted in the characterization of an atropopeptide that is more extensively modified than previously identified members, resulting in an even more rigid 3-dimensional shape. Moreover, one characterized atropopeptide BGC encoding a single P450 is involved in the biosynthesis of two peptides with the same sequence but distinct and non-overlapping modification patterns. This work expands the atropopeptide chemical space, advances our understanding of atropopeptide biosynthesis and underscores the potential of machine learning in uncovering the uncharted biosynthetic diversity encoded in RiPP biosynthetic blueprints.