Identification of novel broad host-range promoter sequences functional in diverse Pseudomonadota by a promoter-trap approach
Identification of novel broad host-range promoter sequences functional in diverse Pseudomonadota by a promoter-trap approach
Roldan, D. M.; Amarelle, V.
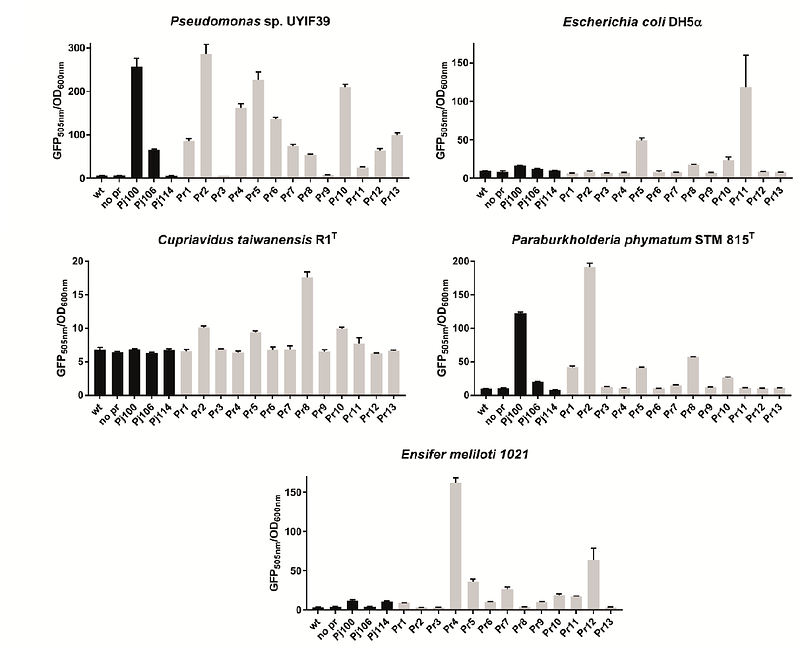
AbstractFinding novel promoter sequences is a cornerstone of synthetic biology. To contribute to the expanding catalog of biological parts, we employed a promoter-trap approach to identify novel sequences within an Antarctic microbial community that act as broad host-range promoters functional in diverse Pseudomonadota. Using Pseudomonas putida KT2440 as host, we generated a library comprising approximately 2,000 clones resulting in the identification of thirteen functional promoter sequences, thereby expanding the genetic toolkit available for this chassis. Some of the discovered promoter sequences prove to be broad host-range as they drove gene expression not only in P. putida KT2440 but also in Escherichia coli DH5, Cupriavidus taiwanensis R1T, Paraburkholderia phymatum STM 815T, Ensifer meliloti 1021, and an indigenous Antarctic bacterium, Pseudomonas sp. UYIF39. Our findings enrich the existing catalog of biological parts, offering a repertoire of broad host-range promoter sequences that exhibit functionality across diverse members of the phylum Pseudomonadota, proving Antarctic microbial community as a valuable resource for prospecting new biological parts for synthetic biology.


