DNA methylation variability defines a fundamental dimension of tumor epigenomes linked to genomic instability, tumor aggressiveness, and clinical outcomes
DNA methylation variability defines a fundamental dimension of tumor epigenomes linked to genomic instability, tumor aggressiveness, and clinical outcomes
Bukovec, D.; Gjorgjioski, B.; Misheva, M. S.; Kungulovski, G.
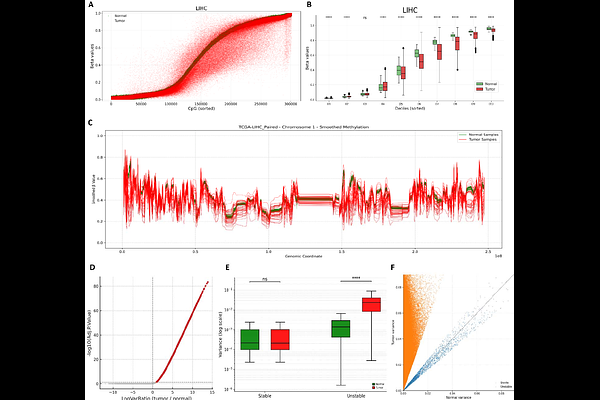
AbstractBackground Tumors exhibit substantial cellular and molecular diversity driven by genetic and epigenetic mechanisms. Large scale profiling efforts have established aberrant DNA methylation as a universal hallmark of cancer. Beyond changes in mean methylation levels, tumor tissues exhibit elevated DNA methylation variability at specific genomic regions within and across tumors. This constitutes a fundamental dimension of cancer epigenomes, reflecting disrupted maintenance of epigenomic states and stochastic drift, which may enable adaptation to the microenvironment, phenotypic plasticity, invasion, disease progression, and treatment resistance. However, the genome wide organization and functional consequences of DNA methylation variability across cancer types remain incompletely understood. Methods We analyzed paired tumor and normal DNA methylation profiles across 16 cancer types to systematically quantify DNA methylation variability. Pan cancer DNA methylation variability was consistently observed using complementary statistical approaches and multiple modes of data representation. We identified cancer specific and pan cancer differentially variable regions and evaluated their associations with genomic features, transcriptional and chromatin regulators, and biological processes. Variability was quantified using three measures per sample: the proportion of intermediately methylated sites (PIM), genome wide Shannon entropy, and a DNA methylation based stemness index. Associations with genomic instability, tumor biological features, and clinical outcomes were subsequently assessed. Results Tumor samples consistently exhibited higher DNA methylation variability than matched normal tissues, reflected by increased dispersion and wider interquartile ranges. Pan cancer variably methylated regions were depleted in promoters and enriched in open sea regions, in heterochromatic H3K27me3 decorated PRC2 repressed domains, and at enhancers. They preferentially contained motifs for transcription factors involved in developmental regulation. Elevated DNA methylation variability, captured by higher PIM, entropy, and stemness scores, was associated with increased genomic instability manifested by higher aneuploidy, increased DNA break points, a greater fraction of the genome altered, and increased tumor mutational burden, as well as with aggressive tumor features such as lymph node involvement, post therapy neoplasm events, and elevated hypoxia scores. Importantly, tumors with high DNA methylation variability exhibited significantly worse overall, progression free, and disease free survival. Conclusions DNA methylation variability is a pervasive and clinically relevant feature of tumor epigenomes, reflecting epigenetic and genetic instability, expanded regulatory plasticity, and tumor aggressiveness.