Coding sequence clustering universally predicts fine- and coarse-scale chromatin compartment landscapes
Coding sequence clustering universally predicts fine- and coarse-scale chromatin compartment landscapes
Cerbus, R. T.; Kawaguchi, K.; Hiratani, I.
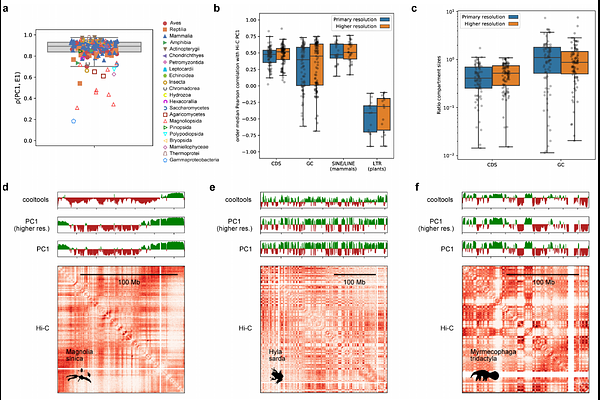
AbstractIntra-chromosomal contact maps from many species display a striking plaid pattern, reflecting chromatin compartmentalization at a sub-chromosomal scale. Although widely regarded as a core feature of genome architecture, such patterns are absent in many organisms, and their underlying determinants remain unclear. Here we systematically examine the relationships among sequence features, chromatin compartmentalization, and evolutionary conservation across 247 species spanning five kingdoms. By testing the usual genomic suspects as determinants of compartmentalization within and between species, we identify the coding DNA sequence (CDS) density landscape as the most consistent predictor of compartmentalization, including whether an organism exhibits a fine-scale plaid or coarse-scale non-plaid chromatin contact map. In contrast, correlations with GC content, CpG density, or repeat element composition vary across lineages, contextualizing long-standing observations such as the prominence of chromosome G-banding in amniotes and its absence elsewhere. Notably, compartment organization is conserved across syntenic blocks between species separated by up to 1 billion years of evolution, and this conservation tracks with preservation of CDS density profiles rather than other sequence features. These findings establish the genomic distribution of coding sequences as a universal and deeply conserved organizing principle of nuclear compartment architecture.