eTRex Reveals Oncogenic Transcriptional Regulatory Programs Across Human Cancers
eTRex Reveals Oncogenic Transcriptional Regulatory Programs Across Human Cancers
Lu, Z.; Yang, Y.; Zheng, Q.; Gao, F.; Xu, L.; Wang, X.
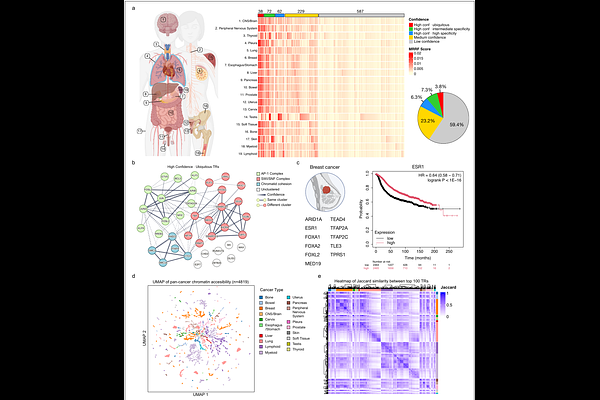
AbstractTranscriptional regulators (TRs) are essential proteins that regulate gene transcription, and their dysregulation defines the oncogenic transcriptional regulatory programs that drive tumor-specific gene expression. Existing pan-cancer resources summarize these programs by aggregating signals across individual datasets, neglecting the context-specific features that capture the diversity of transcriptional regulation underlying cancer heterogeneity. By developing a variational Bayesian hierarchical model named eTRex (epigenomics-based Transcriptional Regulator explorer) and applying it to 4,819 cancer-related ATAC-seq datasets, we provide a comprehensive pan-cancer atlas of functional TR profiles that preserve the context-specific features of each dataset. This study reveals both common regulators across diverse malignancies and those with highly specific roles. We extensively validated these findings using independent CRISPR/Cas9 screening, mutation, and transcriptomic datasets. Collectively, this pan-cancer atlas of functional TR profiles represents a comprehensive, biologically interpretable resource for uncovering transcriptional regulatory programs, identifying biomarkers, prioritizing therapeutic targets in oncology, and is freely accessible through an interactive web portal.