BiGAT-Fusion: Node-Wise Gated Bidirectional Graph Attention for Drug Repurposing
BiGAT-Fusion: Node-Wise Gated Bidirectional Graph Attention for Drug Repurposing
Ding, W.
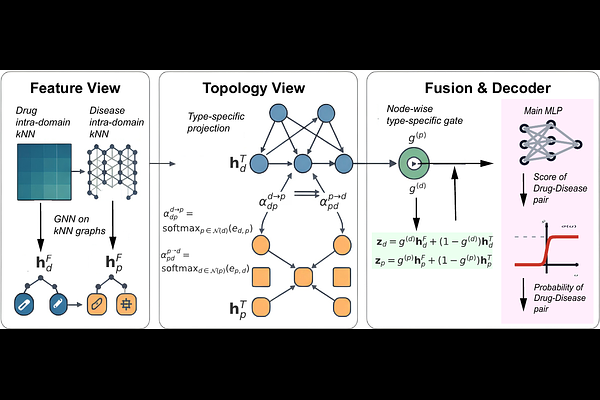
AbstractDrug discovery remains slow, costly, and failure-prone, motivating computational drug repurposing that prioritizes plausible drug-disease associations (DDAs). However, DDA prediction faces three stubborn challenges: (i) extreme class imbalance with few confirmed links amid vast unknowns; (ii) directional asymmetry on the bipartite drug-disease graph that standard message passing underutilizes; and (iii) static fusion of heterogeneous evidence, where fixed rules cannot adapt the relative value of feature (similarity) and topology (association) views across nodes. We present BiGAT-Fusion, a two view graph neural model that addresses these issues end-to-end. Feature view embeddings are learned on kNN graphs built from drug-drug and disease-disease similarities, while topology view embeddings are learned via a bidirectional graph attention layer that explicitly models drug-to-disease and disease-to-drug aggregation. The views are combined by type specific, node wise gates that adaptively weight evidence per node. Pair scores are produced by a residual mixture of experts head that augments a main Multi-Layer Perceptron (MLP) with a bounded low rank bilinear residual and node biases. Under repeated K-fold cross validation with validation driven selection, and with the topology graph constructed strictly from training positives, BiGAT-Fusion achieves state-of-the-art AUPRC on standard benchmarks (Gdataset, Cdataset, LRSSL, Ldataset) while remaining competitive in AUROC. Analyses of learned gates and directional attentions corroborate the model's design. BiGAT-Fusion thus offers a practical, interpretable component for large-scale, computer-aided drug repurposing.