Linking genotype to longevity under genealogical discordance in Sebastes rockfishes
Linking genotype to longevity under genealogical discordance in Sebastes rockfishes
Mo, Y. K.; Sudmant, P. H.; Hahn, M. W.
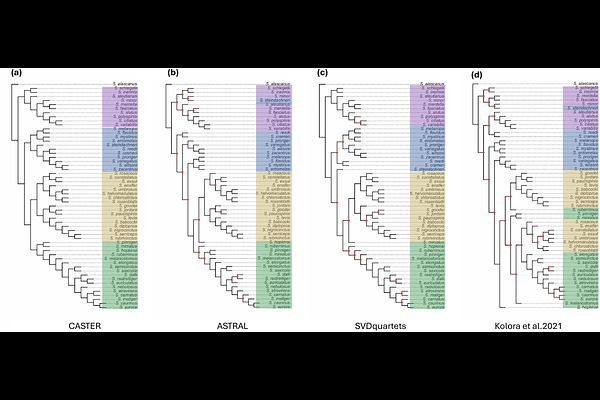
AbstractRockfishes (genus Sebastes) show extreme variation in longevity among closely related species, but the evolutionary history of this young radiation is highly complex. To unpack these relationships and to associate genotypes with phenotypes, we quantified genealogical discordance among 55 Sebastes species and implemented a phyloGWAS framework that incorporates discordant gene histories into genotype-longevity association tests. We found that genealogical discordance is extremely high: the inferred species tree topology differed among several ILS-aware methods, with most internal branches having low concordance factors regardless of which method was used. Nevertheless, some phylogenetic structure was shared by all inferred species trees. We used simulations to assess the statistical properties of phyloGWAS applied to complex traits using different genetic relatedness matrices (GRMs) and under varying levels of discordance. Adding an accurate GRM reduced false positives relative to a model without relatedness, but GRMs only modestly increased power to detect true positives. Using multiple approaches on the Sebastes data, phyloGWAS identified several variants associated with longevity. Our results indicate that extreme genealogical discordance is a core feature of Sebastes evolution and that phyloGWAS can help in connecting genotype to phenotype under these conditions.