Native mass spectrometry of membrane protein-lipid interactions in different detergent environments
Native mass spectrometry of membrane protein-lipid interactions in different detergent environments
Kumar, S.; Stover, L.; Wang, L.; Bahramimoghaddam, H.; Zhou, M.; Russll, D. H.; Laganowsky, A.
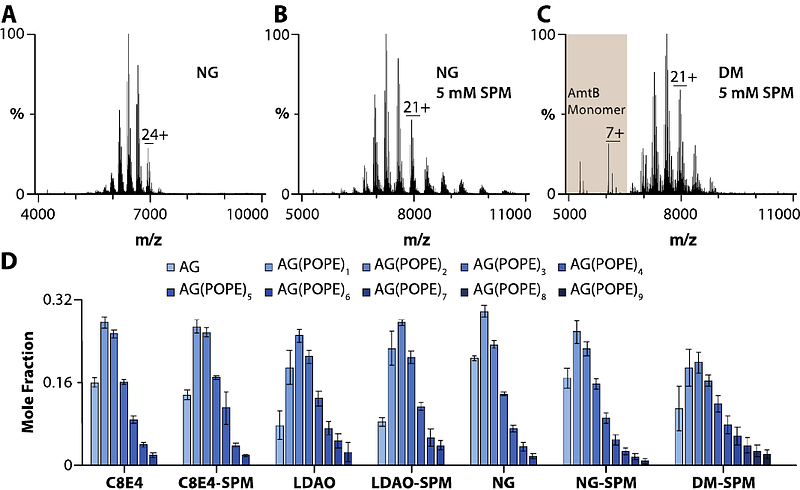
AbstractNative mass spectrometry (MS) is revealing the role of specific lipids in modulating membrane protein structure and function. Membrane proteins solubilized in detergents are often introduced into the mass spectrometer; however, commonly used detergents for structural studies, such as dodecylmaltoside, tend to generate highly charged ions, leading to protein unfolding, thereby diminishing their utility for characterizing protein-lipid interactions. Thus, there is a critical need to develop approaches to investigate protein-lipid interactions in different detergents. Here, we demonstrate how charge-reducing molecules, such as spermine and trimethylamine-N-oxide, enable characterization of lipid binding to the bacterial water channel (AqpZ) and ammonia channel (AmtB) in complex with regulatory protein GlnK in different detergent environments. We find protein-lipid interactions are not only protein-dependent but can also be influenced by the detergent and type of charge-reducing molecule. AqpZ-lipid interactions are enhanced in LDAO (n-dodecyl-N,N-dimethylamine-N-oxide), whereas the interaction of AmtB-GlnK with lipids is comparable among different detergents. A fluorescent lipid binding assay also shows detergent dependence for AqpZ-lipid interactions, consistent with results from native MS. Taken together, native MS will play a pivotal role in establishing optimal experimental parameters that will be invaluable for various applications, such as drug discovery, as well as biochemical and structural investigations.


