Signal, noise, and bias in phylogenetic inference:potential and limits to the resolution of phylogenetic trees in the phylogenomic era
Signal, noise, and bias in phylogenetic inference:potential and limits to the resolution of phylogenetic trees in the phylogenomic era
Dornburg, A.; Su, Z. T.; Jin, Y.; Fisk, N.; Townsend, J. P.
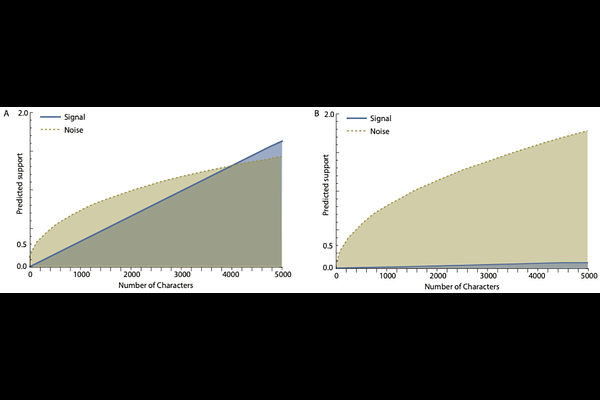
AbstractPhylogenomic datasets assembled to resolve the Tree of Life now routinely span thousands of loci comprising millions of characters. Yet the persistence of incongruent topologies across such datasets reveals a fundamental truth of phylogenetics: not all data are equally informative. Here we derive analytical approaches that predict the relative impacts of phylogenetic signal, stochastic noise, and systematic bias on phylogenetic inference. We show that these three components exhibit divergent scaling properties with character sampling: signal and bias accumulate linearly, while noise accumulates nonlinearly with a concave trajectory. For some phylogenetic problems, substantial amounts of phylogenetic noise may eventually be overwhelmed by signal. For other phylogenetic problems - especially those involving deep divergences, short internodes, or constrained character-state space - the slope of signal accumulation can be so shallow that even signal from genome-scale data may never practically exceed noise. Moreover, linear accumulation of phylogenetic bias can in principle continuously overwhelm accumulation of signal at a lower slope with additional characters, regardless of dataset size. Applying our theory to empirical datasets, we show that anchored hybrid enrichment and ultraconserved element loci, like any loci, can exhibit signal that is overwhelmed by noise, and that character acquisition biases in some loci can further confound inference. Given the pervasive nature of incongruence in the phylogenomic era, our work provides a theoretical foundation for understanding the limits of inference, improving experimental design, and guiding efficient and accurate resolution of the Tree of Life.