Multi-omics analysis and genome-scale metabolic reconstruction of cattle Bos taurus for optimal production of cultured meat
Multi-omics analysis and genome-scale metabolic reconstruction of cattle Bos taurus for optimal production of cultured meat
Lee, J.; Kim, J.; Bae, H. W.; Kim, M.; Jung, B. K.; Kim, J.; Lee, S.; Kim, H. U.
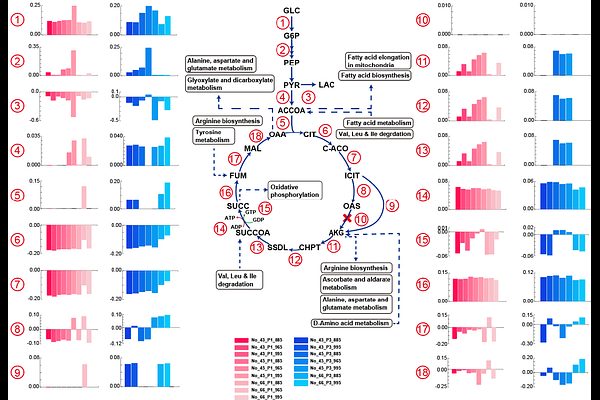
AbstractWith the growing urgency of addressing climate change, cultured meat has gained significant attention as a sustainable alternative to conventional meat production. Bos taurus, a key cattle species, is considered as a potential source of cultured meat. However, much remains to be understood about the biology of B. taurus muscle cells. In this study, bovine satellite cells (BSCs) derived from the semimembranosus muscle of Korean Hanwoo cattle were subjected to multi-omics profiling and genome-scale metabolic reconstruction. First, differential gene expression and gene set enrichment analyses, based on RNA-seq data, identified key pathways associated with muscle cell proliferation (e.g., \'Cell cycle and \'RNA polymerase\') and differentiation (e.g., \'Cytoskeleton in muscle cells\' and \'Tryptophan metabolism\'). Next, using the human1 GEM as a template, we constructed the first B. taurus-specific genome-scale metabolic model (GEM), named BtaSBML2986, which comprises 2,986 genes, 13,278 reactions, and 8,652 metabolites. Muscle cells were cultured under six distinct conditions, and biomass predictions generated using BtaSBML2986 were validated against experimental growth rates. This integrated approach also provided insights into core pathways such as glycolysis and the TCA cycle. BtaSBML2986 represents a significant step forward in understanding B. taurus muscle metabolism and will serve as a valuable tool for advancing cultured meat research and optimizing culture processes.