Structural mechanism of HP1-dependent transcriptional repression and chromatin compaction
Structural mechanism of HP1-dependent transcriptional repression and chromatin compaction
Sokolova, V.; Miratsky, J.; Svetlov, V.; Brenowitz, M.; Vant, J.; Lewis, T. S.; Dryden, K.; Lee, G. L.; Sarkar, S. L.; Nudler, E.; Singharoy, A.; Tan, D.
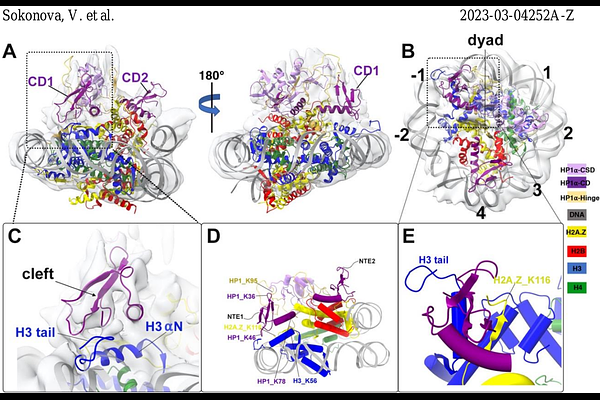
AbstractHeterochromatin protein 1 (HP1) plays a central role in establishing and maintaining constitutive heterochromatin. However, the mechanisms underlying HP1-nucleosome interactions and their contributions to heterochromatin functions remain elusive. In this study, we employed a multidisciplinary approach to unravel the interactions between human HP1 and nucleosomes. We have elucidated the cryo-EM structure of an HP1 dimer bound to an H2A.Z nucleosome, revealing that the HP1 dimer interfaces with nucleosomes at two distinct sites. The primary binding site is located at the N-terminus of histone H3, specifically at the trimethylated K9 (K9me3) region, while a novel secondary binding site is situated near histone H2B, close to nucleosome superhelical location 4 (SHL4). Our biochemical data further demonstrates that HP1 binding influences the dynamics of DNA on the nucleosome. It promotes DNA unwrapping near the nucleosome entry and exit sites while concurrently restricting DNA accessibility in the vicinity of SHL4. This study offers a model that explains how HP1 functions in heterochromatin maintenance and gene silencing, particularly in the context of H3K9me-dependent mechanisms. Additionally, it sheds light on the H3K9me-independent role of HP1 in responding to DNA damage.