Insights into tick-pathogen interactions - a single cell RNA sequencing approach of transcriptional changes during ehrlichial infection
Insights into tick-pathogen interactions - a single cell RNA sequencing approach of transcriptional changes during ehrlichial infection
Adegoke, A.; Aspinwall, J.; McNinch, C.; Ho, M.; Miranda, A. X.; Hoyt, F. H.; Nair, V.; Lack, J.; Saito, T. B.
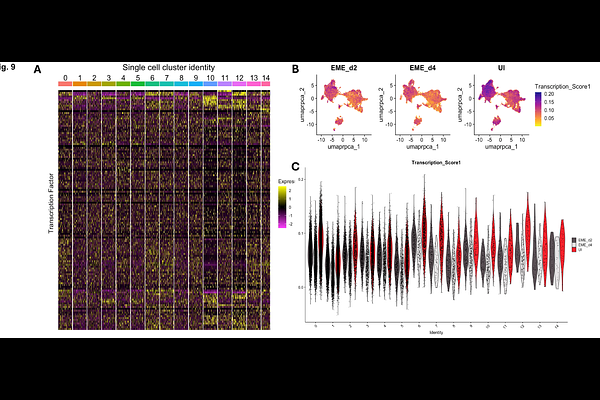
AbstractTick-borne diseases represent a significant threat to human and animal health worldwide. In the United States, the blacklegged tick, Ixodes scapularis (I. scapularis), serves as a competent vector for several bacterial pathogens, including Ehrlichia muris eauclairensis (EME). The I. scapularis embryonic cell line (ISE6) is a valuable tool for propagating tick-borne pathogens and studying tick-pathogen interactions. In this study, we examined the cellular complexity of ISE6 cells and their response to EME infection. Single-cell RNA sequencing revealed 15 distinct cell clusters present. Although ISE6 cells are heterogeneous, they do not display transcriptional similarity to any known tick tissues. Notably, this lack of similarity did not influence their susceptibility to EME infection. Our results demonstrated that EME infection induces time-dependent transcriptional changes in ISE6 cells: early infection is characterized by upregulation of genes associated with stress adaptation, mitochondrial function, and metabolic pathways, whereas late infection leads to broad downregulation of genes involved in the cell cycle, DNA replication, and cytoskeletal organization. These findings enhance our understanding of ehrlichial interactions with ISE6 cells and reinforce the utility of this cell line as a resource for isolating and propagating arthropod endosymbionts and tick-borne pathogens.