Nucleotide-level chemical reaction network modeling enables quantitative prediction of reconstituted cell-free expression system
Nucleotide-level chemical reaction network modeling enables quantitative prediction of reconstituted cell-free expression system
Jurado, Z.; Pandey, A.; Murray, R. M.
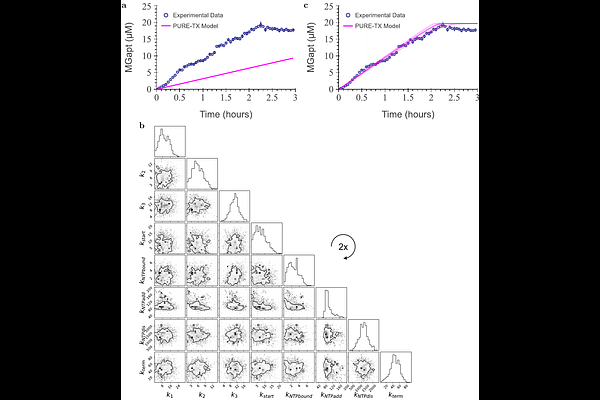
AbstractCell-free expression systems offer a method for rapid prototyping of DNA circuits and functional protein synthesis. While crude extracts remain a black box with many components carrying out unknown reactions, PURE contains only the required transcription and translation components for protein production. All proteins and small molecules are at known concentrations, enabling detailed modeling for reliable computational predictions. However, there is little to no experimental data supporting the expression of target proteins for PURE-based models. In this work, we generalized the PURE detailed translation model for proteins with arbitrary amino acid compositions and lengths. We then built a chemical reaction network for transcription in PURE, validating the transcription models using DNA expression for the malachite-green aptamer (MGapt) to measure mRNA production. Lastly, we coupled the transcription and the generalized translation models to create a PURE protein synthesis model built purely of mass-action reactions. We used the combined model to capture the kinetics of MGapt and deGFP expressed from plasmids at varying concentrations.