Targeted DNA methylation editing in vivo
Targeted DNA methylation editing in vivo
Kalomoiri, M.; Sorini, C.; Vos, S. V. T.; Camargo, A.; Prakash, C. R.; Svenningsson, P.; Pahlevan Kakhki, M.; Kular, L.; Jagodic, M.
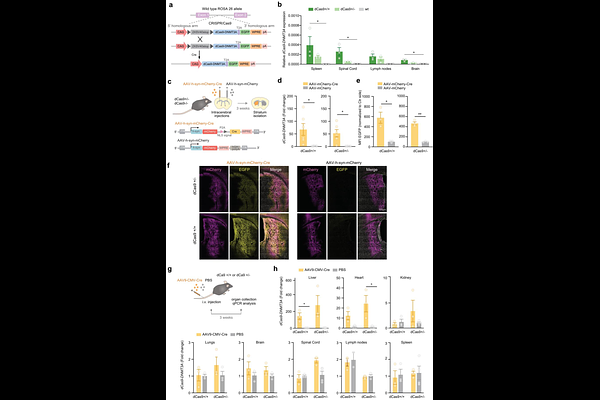
AbstractThe number of epigenome-wide association studies linking CpG DNA methylation with disease, traits and exposures, continues to rise. Establishing the causation of these associations, despite the rapid development of epigenome editing tools, remains challenging, particularly in vivo. In this study, we developed and characterized three Cre-dependent CRISPR-based mouse lines that enable locus-specific DNA methylation deposition by either constitutive or inducible dCas9-DNMT3A expression. We demonstrate robust locus-specific DNA methylation deposition of MHC class II (H2-Ab1) and interleukin 6 (Il6) genes in bone marrow-derived myeloid cells ex vivo. Moreover, neuron-specific targeting results in decreased cannabinoid receptor 1 (Cnr1) expression in striatal neurons in vivo. Notably, we demonstrate that the causal effect of DNA methylation on gene expression is locus-dependent, reinforcing the necessity of such editing tools for detailed understanding of the role of DNA methylation and for addressing the causality of disease-associated CpGs.