Systematics, diversification, and biogeography of Macromiidae (Odonata: Anisoptera)
Systematics, diversification, and biogeography of Macromiidae (Odonata: Anisoptera)
Uche Dike, R.
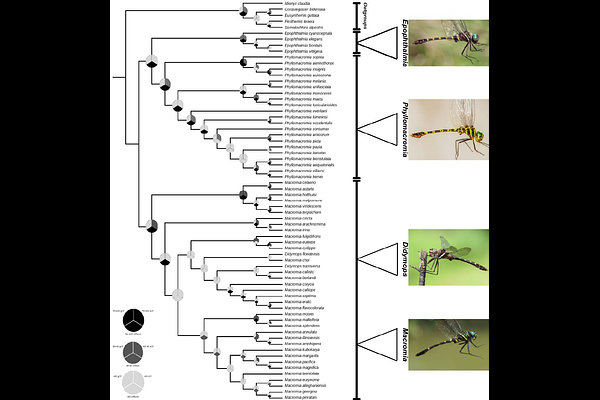
AbstractMacromiidae is a widely distributed lineage of libelluloid dragonflies with a largely allopatric genus-level distribution across the Holarctic, Afrotropical, Australasian, and Indo-Malayan regions. Previous studies involving this family have been complicated by morphological convergence and limited phylogenetic sampling. Here, we present the most densely sampled phylogenetic framework for Macromiidae to date, using Anchored Hybrid Enrichment data from 62 of the 125 described species. Our sampling represents all four genera and major geographic regions, including Libelluloid and Cordulegastrid outgroups. Maximum likelihood recovered three major lineages: Epophthalmia, Phyllomacromia, and Macromia sensu lato, with Epophthalmia strongly supported as sister to Phyllomacromia. Didymops was not recovered as monophyletic and was placed within Macromia, although deeper relationships within the Macromia complex showed some gene tree discordance. We additionally scored seven male genitalic characters and reconstructed their evolution across a dated phylogeny. We revealed that these traits varied heavily in phylogenetic signal, with some characters supporting the major clades and others showing high degree of homoplasy. Fossil-calibrated divergence time estimation placed the crown origin of Macromiidae in the late Oligocene (24 Ma), with other major intrafamilial divergences concentrated in the Miocene. Historical biogeographic reconstructions consistently supported Afrotropical origins for Phyllomacromia, Indo-Malayan centered ancestry for Epophthalmia, and a multi-region history for Macromia + Didymops spanning Indo-Malayan, Australasian, and Nearctic regions. Habitat reconstructions favored lentic ancestry for Macromiidae, and diversification rate variation was best explained by trait-independent models rather than lentic/lotic habitat association.