STCS: A Platform-Agnostic Framework for Cell-Level Reconstruction in Sequencing-Based Spatial Transcriptomics
STCS: A Platform-Agnostic Framework for Cell-Level Reconstruction in Sequencing-Based Spatial Transcriptomics
Chen Wu, L.; Hu, X.; Zhan, F.; Sun, C.; Gonzales, J.; Ofer, R.; Tran, T.; Verzi, M. P.; Liu, L.; Yang, J.
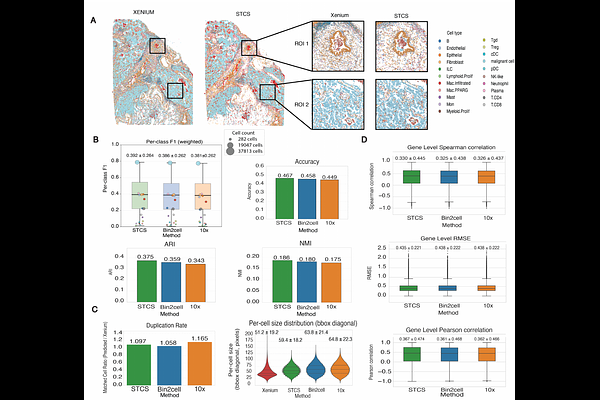
AbstractSequencing-based spatial transcriptomics platforms such as Visium HD and Stereo-seq achieve transcriptome-wide coverage at subcellular resolution, yet their measurements are defined over spatially barcoded units rather than biologically segmented cells. Reconstructing coherent cell-level expression profiles from these data remains a central computational challenge. Here, we introduce Spatial Transcriptomics Cell Segmentation (STCS), a platform-agnostic framework that reconstructs single-cell expression profiles by assigning spatial units to nuclei segmented from paired H&E images, using a combined transcriptomic and spatial distance. STCS is governed by two interpretable parameters that can be selected using reference-free internal metrics. On both Visium HD human lung cancer data with matched Xenium references and Stereo-seq mouse brain data, STCS achieves consistent improvements over existing methods across multiple evaluation dimensions. STCS is fully open-source and designed for broad applicability across sequencing-based spatial transcriptomics technologies.