Leveraging protein language and structural modelsfor early prediction of antibodies with fast clearance
Leveraging protein language and structural modelsfor early prediction of antibodies with fast clearance
Ramanujan, S.; Mazrooei, P.; O'Neil, D.; Chen, B.; Izadi, S.
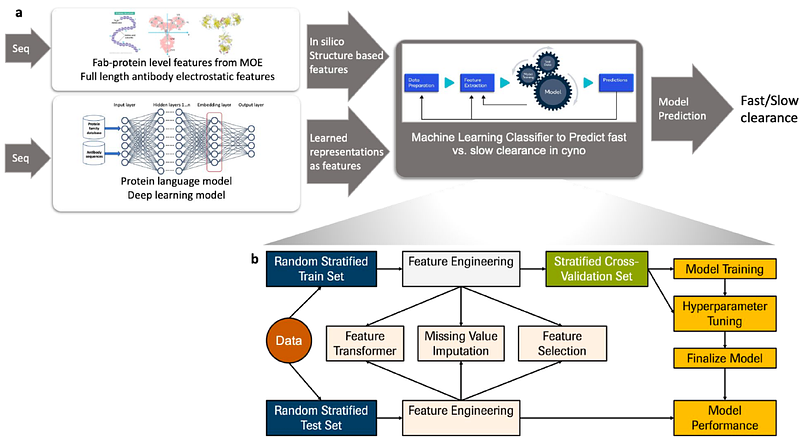
AbstractMonoclonal antibodies (mAbs) with long systemic persistence are widely used as therapeutics. However, antibodies with atypically fast clearance require more dosing, limiting their clinical usefulness. Deep learning can facilitate using sequence-based modeling to predict potential pharmacokinetic (PK) liabilities before antibody generation. Assembling a dataset of 103 mAbs with measured nonspecific clearance in cynomolgus monkeys (cyno), and using transfer learning from large protein language models, we developed multiple machine learning models to predict mAb clearance as fast/slow clearing. Focusing on minimizing misclassification of potentially promising molecules as fast clearing, our results show that using physicochemical properties yielded up to 73.1+/-1.1% classification accuracy on hold-out test data (precision 65.2+/-2.3%). Using only sequence-based features from deep learning protein language models yielded a comparable performance of 71+/-1.4% (precision 65.5+/-2.5%). Combining structural and deep learning derived features yielded a similar accuracy of 73.9+/-1.1%, and slightly improved precision (68.3+/-2.4%). Features important for classifying fast/slow clearance point to charge, moment, and surface area properties at pH 7.4 as well as deep learning derived features. These results suggest that the protein language models provide comparable information and predictive performance of clearance as physicochemical features. This work provides a foundation for in silico prediction of protein pharmacokinetics to inform antibody candidate generation and early deprioritization of designs with high risk of fast clearance. More generally, it illustrates the value of transfer learning-based application of protein language models to address characteristics of importance for protein therapeutics.


