A harmonized benchmarking framework for implementation-aware evaluation of 46 polygenic risk score tools across binary and continuous phenotypes
A harmonized benchmarking framework for implementation-aware evaluation of 46 polygenic risk score tools across binary and continuous phenotypes
Muneeb, M.; Ascher, D.
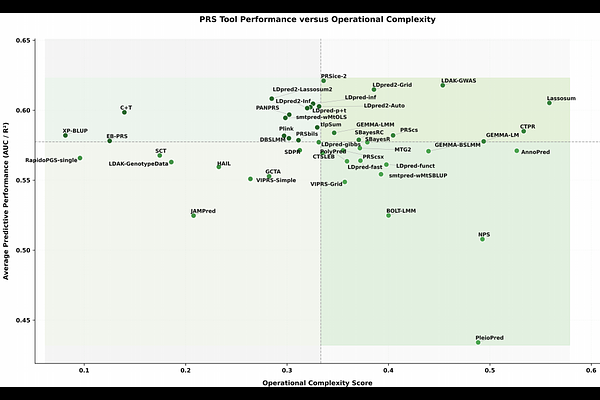
AbstractPolygenic risk score (PRS) tools differ substantially in statistical assumptions, input requirements, and implementation complexity, making direct comparison difficult. We developed a harmonized, implementation-aware benchmarking framework to evaluate 46 PRS tools across seven binary UK Biobank phenotypes and one continuous trait under three model configurations: null, PRS-only, and PRS plus covariates. The framework integrates standardized preprocessing, tool-specific execution, hyperparameter exploration, and unified downstream evaluation using five-fold cross-validation on high-performance computing infrastructure. In addition to predictive performance, we assessed runtime, memory use, input dependencies, and failure modes. A Friedman test across 40 phenotype--fold combinations confirmed significant differences in tool rankings ({chi}2 = 102.29, p = 2.57 x 10-11), with no single method universally optimal. These findings provide a reproducible framework for comparative PRS evaluation and demonstrate that tool performance is shaped not only by statistical methodology but also by phenotype architecture, preprocessing choices, covariate structure, computational demands, software robustness, and practical implementation constraints.