Geometry-enhanced protein language modeling enables discovery of novel antibiotic resistance genes
Geometry-enhanced protein language modeling enables discovery of novel antibiotic resistance genes
Lin, X.; Guan, J.; Hong, Y.; Guo, Y.; Yang, Y.; Xie, P.; Zhao, Z.; Liu, X.; Huang, Y.; Ye, Y.; Tang, Y.; Lee, T.-Y.; Chiang, Y.-C.; Wei, L.; Liu, X.; Wang, J.; Pan, Y.; Tang, J.; Pei, Y.; Yao, L.
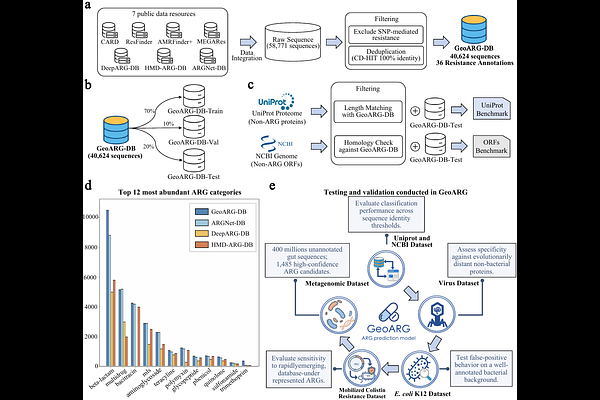
AbstractThe global antibiotic resistome remains largely unexplored, not because antibiotic resistance genes (ARGs) are rare in the environment, but because many are evolutionarily distant from known ARGs. Current computational approaches primarily rely on sequence homology, and thus miss distant homologues. We develop GeoARG, a geometry-enhanced framework that integrates structural features with protein language models through knowledge distillation, enabling efficient large-scale screening using sequence input alone. Across multiple benchmarks, GeoARG substantially improves the detection of remotely homologous ARGs, particularly under low sequence identity and fragmented conditions. Large-scale metagenomic analysis uncovers 1,485 high-confidence ARG candidates that are highly divergent from known ARGs, expanding the phylogenetic and functional landscape of the resistome. Structural analyses further show that these candidates preserve active-site geometry and maintain stable ligand-binding configurations consistent with known resistance mechanisms. These results demonstrate that geometric constraints enable systematic expansion of the resistome and facilitate the discovery of evolutionarily distant yet functionally conserved genes. A public web server is available at https://ycclab.cuhk.edu.cn/GeoARG/