In-situ Target Base Editing Combining with Biosensor-driven Strategy Reveals Critical Single Nucleotide Variants for Enhanced Recombinant Protein Secretion in Pichia pastoris
In-situ Target Base Editing Combining with Biosensor-driven Strategy Reveals Critical Single Nucleotide Variants for Enhanced Recombinant Protein Secretion in Pichia pastoris
Tang, Y.; Zhang, C.
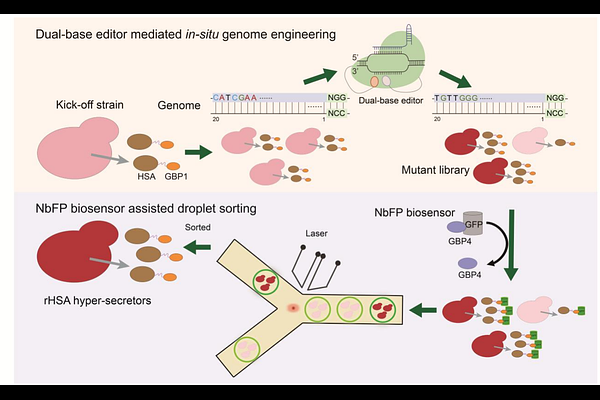
AbstractThe disparity between the production and demand of recombinant proteins (r-proteins) has significantly hindered their commercial viability. Leveraging genomic resources offers substantial promise in enhancing our comprehension of metabolic and regulatory networks, thus facilitating the development of highly productive protein cell factories. However, the considerable gap between high-throughput strategies for monitoring r-protein secretion and genome perturbation in P. pastoris continues to obstruct the systematic linkage of genotype and phenotype, thereby limiting the optimization of production. Here, we developed a novel strategy combining dual-base editor-mediated in-situ genome engineering with nanobody-regulated biosensor-assisted droplet sorting to enhance r-protein secretion (BINDER) in P. pastoris. We successfully employed BINDER to screen recombinant human serum albumin (rHSA) hyper-producers and identified two critical SNVs conferring up to a 1.78-fold improved secretion titer from 113632 mutants, providing valuable insights into the secretion mechanism. Fed-batch cultivation of the engineered strain resulted in the highest reported rHSA titer, 23.43g/L, in P. pastoris, demonstrating its substantial potential for industrial applications. Given the high transferability of base editors and the novel biosensor's independence from the properties of the target protein, the strategy developed here might be expanded to a variety of microbial species and r-proteins.