Programmable nanobody circuits for cell selection
Programmable nanobody circuits for cell selection
Wang, N. B.; Blanch-Asensio, A.; Cevasco, H.; Ploessl, D. S.; Gumustop, D. R.; Ehmann, M. E.; Castellanos, M. F.; Sanchez-Rivera, F. J.; O'Shea, T. M.; Galloway, K. E.
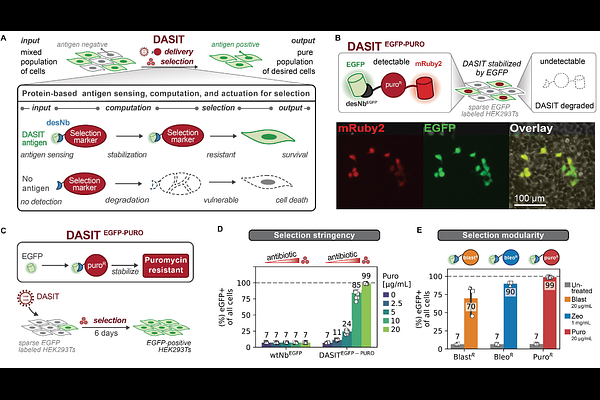
AbstractEfficient and scalable isolation of specific cell populations remains a central bottleneck for genome engineering, pooled screening, and cell therapy manufacturing. Here, we present DASIT (Destabilized-nanobody Antigen Selection and Identification Tool), a protein-based circuit for antigen-specific cell selection. DASIT uses a destabilized nanobody fused to an antibiotic resistance protein. In cells expressing the target antigen, binding of the nanobody fusion to the cognate antigen stabilizes DASIT, thereby coupling the presence of an antigen to a selectable signal. We developed DASIT circuits that enable robust selection of antigen-expressing cells and show that they can be designed to target distinct antigen classes and perform across cell types. Because DASIT operates at the protein level, it supports both stable integration and transient delivery, enabling recyclable selection without permanent genomic integration of resistance markers. We demonstrate scalable, FACS-free enrichment in three challenging applications: multiplexed, logic-gated integration of landing pads in human iPSCs, high-throughput CRISPR screening, and phenotypic selection of in vitro-derived neurons at transplantation scale. By decoupling selection from vector integration, DASIT establishes an automation-compatible architecture for multistep genome engineering, high-throughput library screening and large-scale cell manufacturing.