Low-dimensional representations of genome-scale metabolism
Low-dimensional representations of genome-scale metabolism
Cain, S.; Merzbacher, C.; Oyarzun, D. A.
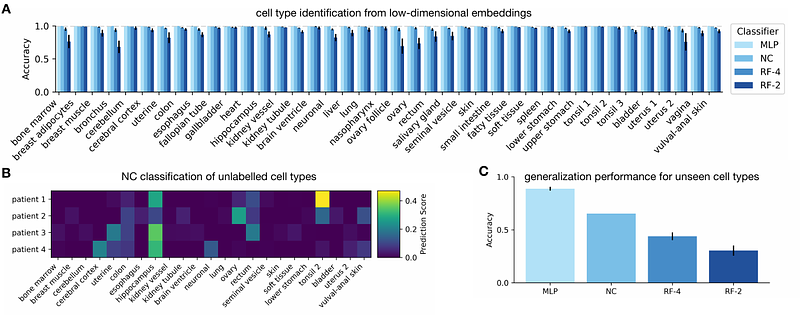
AbstractCellular metabolism is a highly interconnected network with thousands of reactions that convert nutrients into the molecular building blocks of life. Metabolic connectivity varies greatly with cellular context and environmental conditions, and it remains a challenge to compare genome-scale metabolism across cell types because of the high dimensionality of the reaction flux space. Here, we employ self-supervised learning and genome-scale metabolic models to compress the flux space into low-dimensional representations that preserve structure across cell types. We trained variational autoencoders (VAEs) on large fluxomic data (N=800,000) sampled from patient-derived models for various cancer cell types. The VAE embeddings have an improved ability to distinguish cell types than the uncompressed fluxomic data, and sufficient predictive power to classify cell types with high accuracy. We tested the ability of these classifiers to assign cell type identities to unlabelled patient-derived metabolic models not employed during VAE training. We further employed the pre-trained VAE to embed another 38 cell types and trained multilabel classifiers that display promising generalization performance. Our approach distils the metabolic space into a semantically rich vector that can be used as a foundation for predictive modelling, clustering or comparing metabolic capabilities across organisms.


