A Scalable Design for Proximity-Inducing Molecules
A Scalable Design for Proximity-Inducing Molecules
Karaj, E.; Venkatarangan, V.; Sindi, S. H.; Siriwongsup, S.; Lee, C.; Pergu, R.; Vedagopuram, S.; Kailass, K.; Tran, K.; Singh, P.; Singh, S.; Kawai, J.; Fung, J. E.; Tefera, M.; Dhaliwal, R.; Chaudhary, S. K.; Keyes, A.; Sadagopan, A.; Boatner, L.; Shah, N. H.; Fehl, C.; Backus, K. M.; Choudhary, A.
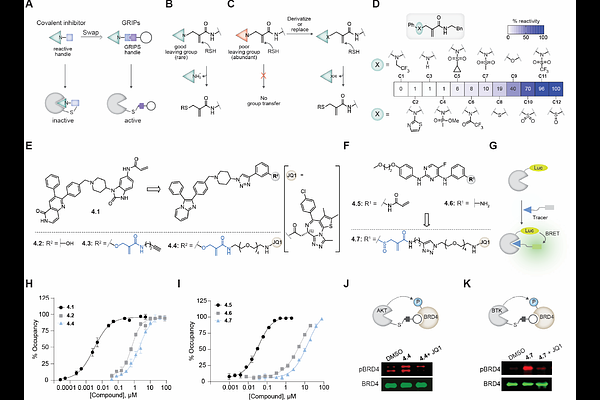
AbstractChimeric molecules, which bring together an effector enzyme and a protein-of-interest (POI) to add/remove post-translational modifications (PTMs), are furnishing transformative modalities (e.g., PROTACs). However, these chimeras' scalability is limited as they employ rare, non-inhibitory binders of effectors. We report GRoup-transfer chimeras for Inducing Proximity (GRIPs) that employ abundantly available effectors' inhibitors to append POI binder on the effector using group-transfer handles. To demonstrate scalability, we develop 6 GRIPs classes for 3 PTMs utilizing diverse inhibitor (16 effector: POI pairs). Furthermore, we report a toolbox of 42 tunable group-transfer handles for Cys/Lys residues and ~5000 inhibitor-residue pairs from diverse effectors. Using global proteomics, we confirm the specificity for group transfer and PTM editing. GRIPs endowed new functionalities to POI drugs, including preventing rebound signaling upon drug withdrawal, a more potent/persistent inhibition, and inhibitor-induced pathway activation in 4 fully-endogenous systems. In diverse hemi-endogenous systems (tagged POI), GRIPs induced condensate formation with reduced off-targets, cleared pathogenic PTMs, and initiated PTM crosstalk. Overall, GRIPs provide a scalable and versatile platform for PTM editing.