High Diversity Gene Libraries Facilitate Machine Learning Guided Exploration of Fluorescent Protein Sequence Space
High Diversity Gene Libraries Facilitate Machine Learning Guided Exploration of Fluorescent Protein Sequence Space
Benabbas, A.; Kearns, P.; Billo, A.; Chisholm, L. O.; Plesa, C.
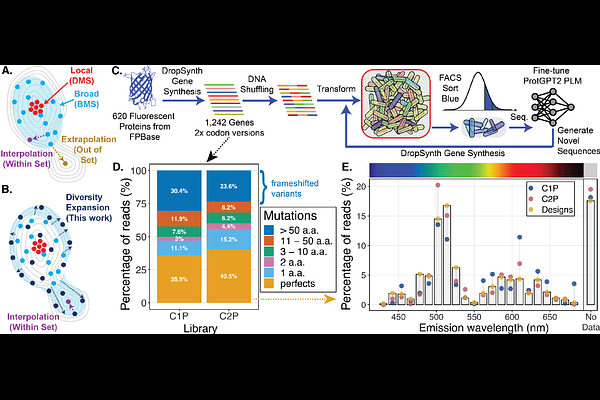
AbstractWhile protein language models (PLMs) have shown great promise for protein design, their performance is fundamentally constrained by the diversity and completeness of available training data. In particular, PLMs often struggle to extrapolate to sequences that fall outside the distribution spanned by their training sets, limiting their ability to discover proteins in sparsely sampled regions of sequence space. Here we test the hypothesis that experimentally expanding training diversity can convert extrapolation into interpolation and thereby enable discovery of functional sequences beyond natural protein manifolds. Using large-scale gene synthesis and DNA shuffling, we generate libraries that span a broad region of fluorescent protein sequence space and create chimeric variants that bridge between distant homologs. Functional screening for blue fluorescence yields thousands of active variants distributed across diverse sequence lineages. Fine-tuning ProtGPT2 on this expanded dataset enables generation of diverse fluorescent proteins, including designs that extend beyond the regions occupied by known natural sequences while retaining function. This work illustrates how synthetic approaches can help address key limitations in machine learning-guided protein design, especially for small or sparsely populated protein families, by actively creating novel sequences across unexplored but functional regions of sequence space.