Molecular Determinants Governing the Antitubercular Activity of Griselimycin
Molecular Determinants Governing the Antitubercular Activity of Griselimycin
Spira, A.; Dash, R.; Lepori, I.; Luo, Y. C.; Newkirk, S.; Bhandari, S.; Siegrist, M. S.; Pires, M.
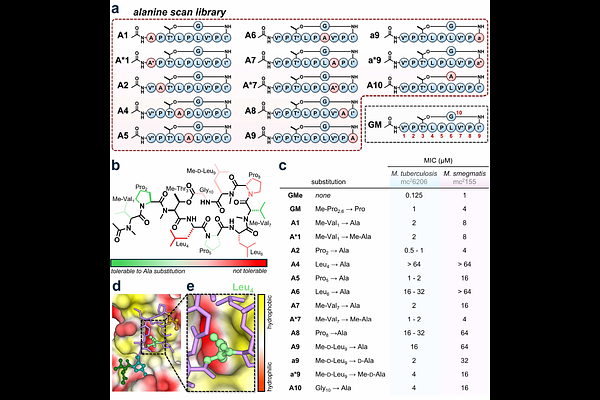
AbstractTuberculosis, often considered the deadliest infectious disease in the world, is associated with over one million deaths annually. The emergence of drug-resistant strains of Mycobacterium tuberculosis (Mtb) makes anti-tuberculosis drug development a critical priority. Griselimycin (GM) is a cyclic peptide that targets the essential DNA sliding clamp of Mtb. While GM is a promising Mtb antibiotic, its poorly understood structure-activity relationship has stalled derivatization. To investigate the contribution of each amino acid towards its activity, we assessed the antibiotic activity of an alanine scan library in M. tuberculosis and M. smegmatis. Residues essential for activity and tolerable to modification were identified, and the impact of backbone N-methylation at each position was determined. Edits to cyclization chemistry, unnatural amino acid incorporation, and replacing the acetylated N-terminus with a free amine were also investigated. Lastly, incorporation of an N-terminal fluorophore enabled visualization of GM accumulation inside of mycobacteria both in and outside of macrophage cells, where Mtb natively resides. These findings present the first comprehensive structure-activity investigation into GM and can be used to rationally design future analogues.