Semantic mining of functional de novo genes from a genomic language model
Semantic mining of functional de novo genes from a genomic language model
Merchant, A. T.; King, S. H.; Nguyen, E.; Hie, B. L.
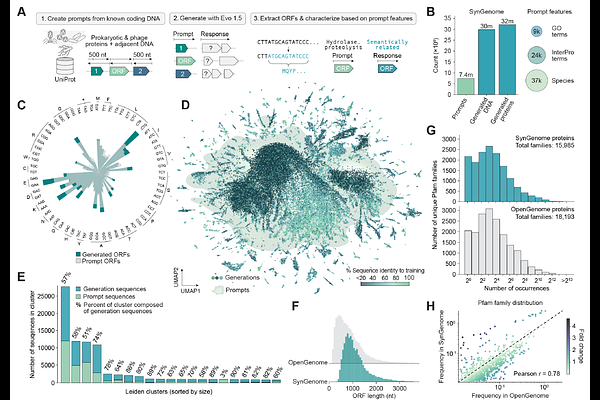
AbstractGenerative genomics models can design increasingly complex biological systems. However, effectively controlling these models to generate novel sequences with desired functions remains a major challenge. Here, we show that Evo, a 7-billion parameter genomic language model, can perform function-guided design that generalizes beyond natural sequences. By learning semantic relationships across multiple genes, Evo enables a genomic \"autocomplete\" in which a DNA prompt encoding a desired function instructs the model to generate novel DNA sequences that can be mined for similar functions. We term this process \"semantic mining,\" which, unlike traditional genome mining, can access a sequence landscape unconstrained by discovered evolutionary innovation. We validate this approach by experimentally testing the activity of generated anti-CRISPR proteins and toxin-antitoxin systems, including de novo genes with no significant homology to any natural protein. Strikingly, in-context protein design with Evo achieves potent activity and high experimental success rates even in the absence of structural hypotheses, known evolutionary conservation, or task-specific fine-tuning. We then use Evo to autocomplete millions of prompts to produce SynGenome, a first-of-its-kind database containing over 120 billion base pairs of AI-generated genomic sequences that enables semantic mining across many possible functions. The semantic mining paradigm enables functional exploration that ventures beyond the observed evolutionary universe.