Sequence-based machine learning reveals 3D genome differences between bonobos and chimpanzees
Sequence-based machine learning reveals 3D genome differences between bonobos and chimpanzees
Brand, C. M.; Kuang, S.; Gilbertson, E. N.; McArthur, E.; Pollard, K. S.; Webster, T. H.; Capra, J. A.
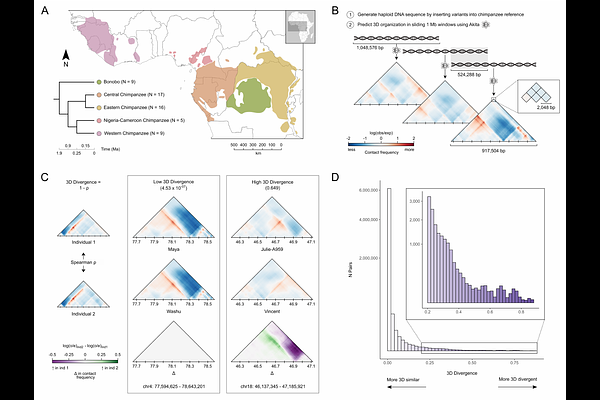
AbstractPhenotypic divergence between closely related species, including bonobos and chimpanzees (genus Pan), is largely driven by variation in gene regulation. The 3D structure of the genome mediates gene expression; however, genome folding differences in Pan are not well understood. Here, we apply machine learning to predict genome-wide 3D genome contact maps from DNA sequence for 56 bonobos and chimpanzees, encompassing all five extant lineages. We use a pairwise approach to estimate 3D divergence between individuals from the resulting contact maps in 4,420 1 Mb genomic windows. While most pairs were similar, ~17% were predicted to be substantially divergent in genome folding. The most dissimilar maps were largely driven by single individuals with rare variants that produce unique 3D genome folding in a region. We also identified 89 genomic windows where bonobo and chimpanzee contact maps substantially diverged, including several windows harboring genes associated with traits implicated in Pan phenotypic divergence. We used in silico mutagenesis to identify 51 3D-modifying variants in these bonobo-chimpanzee divergent windows, finding that 34 or 66.67% induce genome folding changes via CTCF binding motif disruption. Our results reveal 3D genome variation at the population-level and identify genomic regions where changes in 3D folding may contribute to phenotypic differences in our closest living relatives.