Oncogenes and tumor suppressor genes are enriched in stop-loss mutations generating protein extensions
Oncogenes and tumor suppressor genes are enriched in stop-loss mutations generating protein extensions
Boll, L. M.; Martorell, J. A.; Khelghati, N.; Camarena, M. E.; Vianello, C.; Garcia-Soriano, J. C.; Santamaria, E.; Artoleta, I.; Saez-Valle, S.; Perera-Bel, J.; Fortes, P.; Alba, M. M.
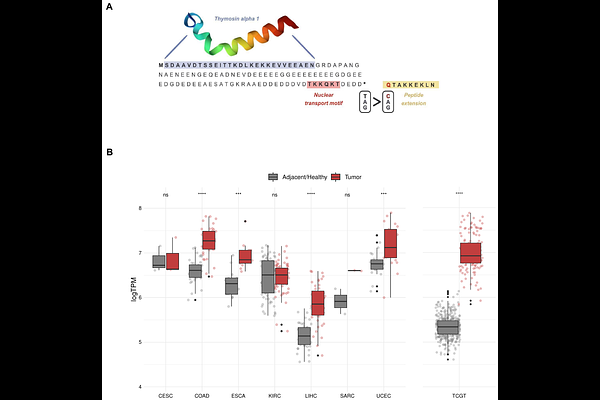
AbstractCancer genomes tend to accumulate a large number of mutations, and even rare mutations such as those causing the loss of a stop codon can be observed in a significant fraction of the tumors. Stop-loss mutations extend protein translation into the 3' untranslated region (3' UTR), generating altered proteins carrying extra amino acid sequences. These C-terminal extensions can potentially have consequences for tumorigenesis and immune recognition. To investigate the prevalence of stop-loss mutations in cancer, and to identify recurrent mutations with a possible tumor-promoting effect, we have interrogated mutation data from the tumor samples of 20,801 patients. This search has resulted in the annotation of 3,757 stop-loss mutations in 3,249 different protein-coding genes. Around 11% of the mutated genes contain recurrent stop-loss mutations, occurring in more than one patient. The protein extensions created by the mutations tend to be hydrophobic and/or positively charged, and these features are associated with an increased propensity to generate MHC I-bound peptides. We have also found that cancer-related genes contain 37% more stop-loss mutations than non-cancer-related genes, with both oncogenes and tumor suppressor genes showing similar enrichments. Furthermore, three out of the four genes with the highest number of stop-loss recurrences, PTMA, PCDH9 and SOX9, are cancer-related. In PTMA, the gene with the largest number of stop-loss mutations (14 patients), the mutation results in an extension of 9 amino acids. We provide experimental evidence that the mutation is associated with impaired cleavage of thymosin alpha 1, a peptide with immunostimulatory functions that is generated from the N-terminal part of the PTMA protein. The study provides evidence that stop-loss mutations are enriched in cancer-associated genes and constitutes a valuable resource for further studies on the effects of stop-loss mutations in cancer.