Genome-wide mapping of gene essentiality in Pseudomonas chlororaphis ATCC 9446 using transposon mutagenesis.
Genome-wide mapping of gene essentiality in Pseudomonas chlororaphis ATCC 9446 using transposon mutagenesis.
De Sandozequi, A.; Bello-Gonzalez, M. A.; Aguilar-Vera, O. A.; Utrilla, J.
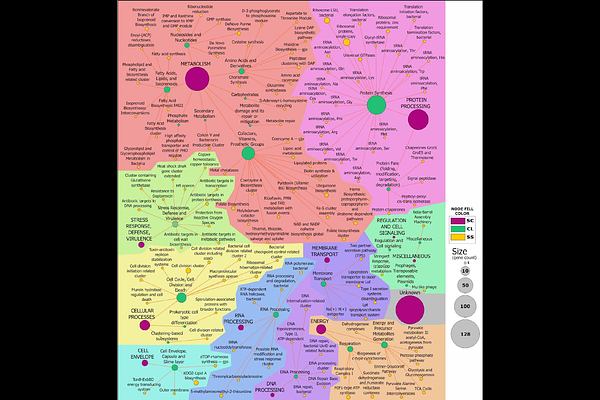
AbstractPseudomonas chlororaphis ATCC 9446 is a non-pathogenic rhizobacterium with biotechnological relevance as a biocontrol agent and a promising chassis for synthetic biology. Understanding which genes are strictly required for survival is fundamental to both bacterial physiology and chassis engineering. Here, we generate a genome-wide map of genetic essentiality for P. chlororaphis using high-density Random Barcoded Transposon Mutagenesis (RB-TnSeq). To convert gene-level annotation into biological insight, we layered functional assignments from complementary annotation pipelines, integrating orthology/domain classifiers, ontology mapping, protein export, and delineation of secondary metabolism. These overlays reveal that essentiality concentrates in canonical information processing, envelope biogenesis, and central energy/cofactor and nucleotide metabolism, while large genomic regions are non-essential and therefore represent safe candidates for streamlining and pathway installation. Mapping essentiality onto biosynthetic gene clusters (BGCs) shows that most pathways are dispensable, but some essential genes co-localize, clarifying boundaries for safe editing around BGC loci. Comparison of experimentally determined essential genes with in silico predictions across additional P. chlororaphis genomes show strong overall agreement. Conversely, 32 essential gene orthogroups were found to be conserved across most genomes, yet were classified as non-essential by a prediction algorithm. Together, the resolved essential genome and its integrative functional interpretation provide a durable reference for P. chlororaphis biology and a functional blueprint that can be leveraged for rational streamlining in agricultural, biocontrol and industrial biotechnology.