Post-fertilization transcription initiation in an ancestral LTR retrotransposon drives lineage-specific genomic imprinting of ZDBF2
Post-fertilization transcription initiation in an ancestral LTR retrotransposon drives lineage-specific genomic imprinting of ZDBF2
Kobayashi, H.; Igaki, T.; Kumamoto, S.; Tanaka, K.; Takashima, T.; Suzuki, S.; Hayashi, M.; Renfree, M. B.; Kawahara, M.; Saito, S.; Kobayashi, T.; Nagashima, H.; Nakano, K.; Matsunari, H.; Uchikura, A.; Kiyonari, H.; Kaneko, M.; Imai, H.; Nakabayashi, K.; Lorincz, M. C.; Kurimoto, K.
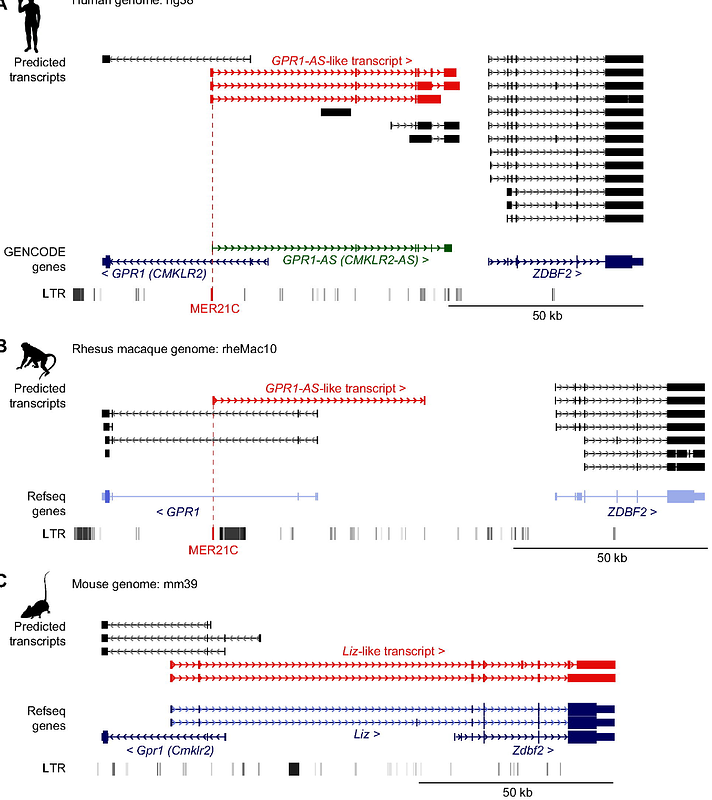
AbstractThe imprinted ZDBF2 gene is controlled by oocyte-derived DNA methylation, but its regulatory system is quite different from that of other canonically imprinted genes that are dependent on DNA methylation deposited in the gametes. At the ZDBF2 locus, maternal DNA methylation in the imprinted differentially methylated region (DMR) does not persist after implantation. Instead, a transient transcript expressed in the early embryo exclusively from the unmethylated paternal allele of the DMR, known as GPR1-AS, contributes to establishing secondary DMRs that maintain paternal expression of ZDBF2 in the somatic lineage. While the imprinting of ZDBF2 and its unique regulatory system are evident in humans and mice, whether this process is conserved in other mammals has not been addressed. Here, we show that the first exon of human GPR1-AS overlaps with that of a long terminal repeat (LTR) belonging to the MER21C subfamily of retrotransposons. Although this LTR family appears and is amplified in eutherians, the MER21C insertion into the GPR1-AS orthologous region occurred specifically in the common ancestor of Euarchontoglires, a clade that includes primates, rodents, and rabbits. Directional RNA sequencing of placental tissues from various mammalian species revealed GPR1-AS orthologs in rabbits and nonhuman primates, with their first exon embedded within the same ancestral LTR. In contrast, allele-specific expression profiling showed that cow and tammar wallaby, mammals outside the Euarchontoglires group, expressed both alleles in all tissues analyzed. Our previous studies showed that LTRs reactivated in oocytes drive lineage-specific imprinting during mammalian evolution. The data presented here suggest that LTR-derived sequence activation after fertilization can also contribute to the lineage-specific establishment of imprinted genes.