A novel proliferative candidate genes panel for idiopathic pulmonary fibrosis: insights from integrated bulk and single-cell RNA sequencing
A novel proliferative candidate genes panel for idiopathic pulmonary fibrosis: insights from integrated bulk and single-cell RNA sequencing
Wang, Q.; Tang, C.; Wu, Q.; Wan, N.; Jin, Z.; yang, C.; Wang, H.; Feng, J.; Wang, Y.
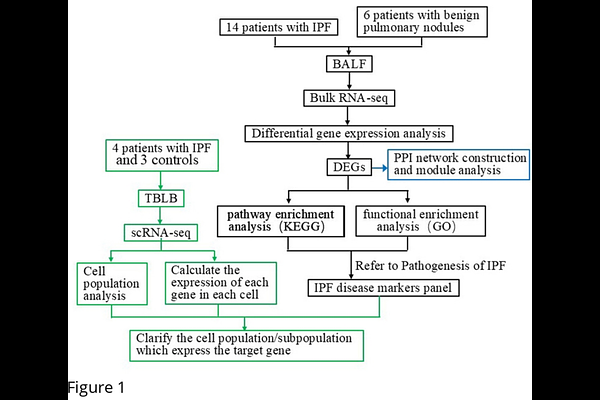
AbstractBackground: Idiopathic pulmonary fibrosis (IPF) remains a fatal interstitial lung disease with limited diagnostic specificity and therapeutic options. This study integrates bulk and single-cell RNA sequencing (RNA-seq) to identify novel biomarkers and elucidate molecular mechanisms underlying IPF pathogenesis. Methods: We prospectively enrolled 14 treatment-naive IPF patients and 6 controls. Bulk RNA-seq was performed on bronchoalveolar lavage fluid (BALF), while single-cell RNA-seq analyzed lung tissues from 4 IPF patients and 3 controls. Differentially expressed genes (DEGs) were identified (|log2FC| >1, FDR <0.05), followed by functional enrichment, protein-protein interaction (PPI) network analysis, and cell-type-specific expression profiling. Results: 1. DEG Identification: Bulk RNA-seq revealed 108 DEGs (24 upregulated, 84 downregulated). KEGG enrichment analysis of DEGs revealed that upregulated genes were mainly enriched in inflammation and immune pathways (such as NF-{kappa}B signaling pathway, Fc epsilon RI signaling pathway, B cell receptor signaling pathway, phagosome, Fc gamma R-mediated phagocytosis), pyrimidine metabolism, cell cycle, and PI3K-Akt signaling pathway. 2. PPI Network: Module analysis identified a proliferative gene module 1 (NUF2, CEP55, ANLN, TTK, TK1, MYBL2, CCNA2, RRM2, CDT1) linked to cell division and cycle regulation. 3. Single-Cell Insights: scRNA-seq of 30,477 cells delineated 11 populations. Module 1 genes exhibited predominant expression in proliferating cells, Module 1 signature score of proliferating cells was significantly higher in IPF than in control group. 4. Pathogenic Links: Key genes (e.g., CEP55, TTK) were associated with PI3K/AKT signaling, epithelial-mesenchymal transition (EMT), and anti-apoptotic pathways, mirroring oncogenic mechanisms. Conclusion: This multi-omics approach uncovers a proliferation-centric gene module in IPF, revealing shared molecular pathways with tumorigenesis. Our findings highlight novel diagnostic biomarkers and suggest repurposing cell cycle inhibitors as potential therapies. Future studies should validate these targets in preclinical models to advance precision medicine for IPF.