Atopic Dermatitis and Psoriasis Differ in Lesional DEG Reference Instability and Non-Lesional Spectrum Displacement: Multi-Cohort Geometric Evidence of Individual Homeostatic Boundary Escape
Atopic Dermatitis and Psoriasis Differ in Lesional DEG Reference Instability and Non-Lesional Spectrum Displacement: Multi-Cohort Geometric Evidence of Individual Homeostatic Boundary Escape
Shabana, B.
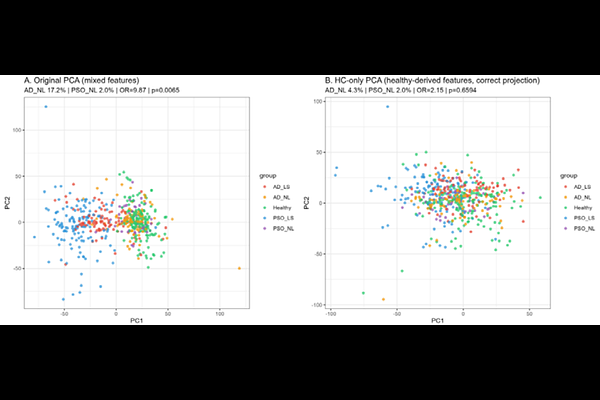
AbstractAtopic dermatitis (AD) transcriptomic studies have produced notoriously inconsistent differential gene expression (DEG) lists across cohorts, and the biological heterogeneity of non-lesional AD skin remains unresolved at the individual level -- limitations that group-averaged DEG analysis is structurally incapable of addressing. We applied a geometric transcriptomic framework to 537 skin RNA-sequencing samples from four independent international cohorts, positioning each sample within a 20-dimensional disease-informative PCA space relative to a unified healthy reference cloud validated by multi-cohort batch integration (ComBat; silhouette width 0.076 across three countries). Applying a bootstrap Jaccard framework to lesional-versus-healthy comparisons, AD lesional DEG signatures were significantly more reference-sensitive than psoriasis (PSO; Jaccard 0.637 vs 0.737; non-overlapping 95% CIs), with per-gene Spearman correlation between absolute effect size and cross-reference reproducibility (rho=0.658, p<2.2x10-16) establishing that AD's instability arises from its structural dependence on smaller-effect-size transcriptomic signals -- a geometric property of AD biology, not a methodological failure of prior studies. In non-lesional skin, spectrum scores -- projections of each sample's displacement from the healthy centroid onto the disease axis -- revealed that AD non-lesional (AD_NL) samples had traversed 22.6% (95% CI 15.2-29.6%) of the healthy-to-lesional axis versus 12.9% (95% CI 8.4-18.2%) for PSO non-lesional (PSO_NL; Wilcoxon p=0.012), with non-lesional skin in each disease displaced toward its own lesional pole at near-identical angles (within-disease delta-angle 1.4 degrees) but AD displaced further. Against a homeostatic boundary defined as the 95th-percentile Euclidean distance of healthy controls from their centroid, 17.2% of AD_NL samples (16/93) individually exceeded this threshold versus 2.0% of PSO_NL (1/49; OR=9.87, p=0.007), replicating across all three cohorts providing AD_NL data. Among boundary-crossing AD_NL samples, two directional patterns emerged: high-positive displacement (n=6) characterised by inflammatory pre-activation (IL-6/JAK-STAT3, IFN-alpha, TNF-alpha/NF-kB) and low/negative displacement (n=10) characterised by broad metabolic suppression and a third transcriptomic axis orthogonal to both canonical disease trajectories -- not attributable to cellular infiltration differences and undetectable by group-level analysis. These findings reframe AD's notorious transcriptomic inconsistency as a predictable consequence of effect-size architecture, establish that a reproducible subset of AD patients harbours individually measurable transcriptomic boundary escape before clinical lesion onset, and identify a biologically uncharacterised non-lesional subgroup that warrants cell-type-resolved investigation as a potential early intervention target.