Altered chromatin accessibility and nucleosome positioning landscape upon HDAC and LSD1 inhibition in cancer cell
Altered chromatin accessibility and nucleosome positioning landscape upon HDAC and LSD1 inhibition in cancer cell
Sen, S.; Esteve, P. O.; Tarasia, D.; Dannenberg, R.; Dey, A.; Maulik, U.; Pradhan, S.; Bandyopadhyay, S.
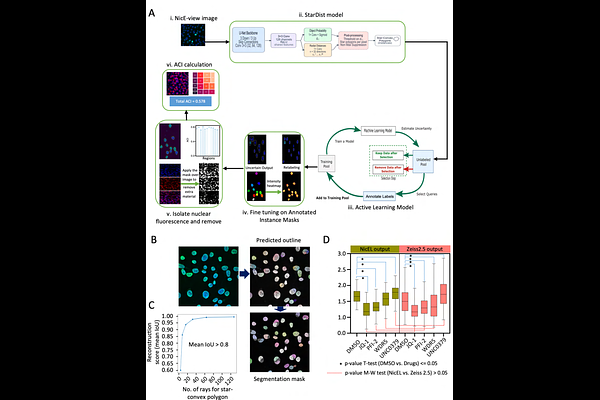
AbstractEpigenetic enzymes, writers, readers and erasers regulate chromatin landscapes and participate in tumor heterogeneity. While therapeutic targeting of these enzymes has shown clinical promise, the comparative efficacy of mono- versus dual-inhibitor strategies remain unclear. Here, we introduce a multi-modal platform that uses NicE-viewSeq and integrates automated deep learning based spatially resolved chromatin accessibility profiling with high-throughput sequencing following epigenetic inhibitor application. Accessible chromatin landscapes were altered along with nucleosome positioning following inhibition of either LSD1 or HDACs alone, or both together. Coordinated modulation of histone marks and the CoREST complex on chromatin was observed across inhibitory conditions. Transcription factor binding analysis identified three predominant families, ETS, RUNT, and bZIP with enhanced chromatin association upon treatments. Mechanistically, a CoREST-RUNX regulatory axis was uncovered wherein JunB, a member of bZIP family displaces CoREST-RUNX at differentially accessible regions, triggering apoptotic pathways. Therefore, JunB-mediated mechanism reveals a convergent therapeutic vulnerability, offering new avenues for optimizing different combinatorial epigenetic therapy in cancer.